micromort
A data package providing curated datasets of micromort (acute risk) and microlife (chronic risk) values from authoritative sources including Wikipedia, CDC MMWR, and academic literature. ## Features
- 62 acute risks measured in micromorts (one-in-a-million death probability per event)
- 35+ chronic factors measured in microlives (30-minute life expectancy change per day)
- 14 authoritative sources with full provenance tracking
- Parquet datasets for cross-language compatibility (R, Python, Arrow)
- Plumber REST API for programmatic access
- Interactive dashboard for data exploration
- targets pipeline for reproducible data updates
Architecture
%%{init: {'theme': 'dark', 'themeVariables': {'primaryColor': '#999999', 'primaryTextColor': '#000000', 'primaryBorderColor': '#CC0000', 'lineColor': '#CC0000', 'secondaryColor': '#999999', 'tertiaryColor': '#999999', 'background': '#000000', 'mainBkg': '#999999', 'nodeBorder': '#CC0000', 'clusterBkg': '#333333', 'clusterBorder': '#CC0000', 'titleColor': '#000000', 'edgeLabelBackground': '#999999'}}}%%
graph LR
Conversion["Unit Conversion<br>5 functions"]
Data["Risk Datasets<br>21 functions"]
Analysis["Risk Analysis<br>6 functions"]
Viz["Visualization<br>5 functions"]
Apps["Interactive Apps<br>6 functions"]
Conversion --> Data --> Analysis --> Viz --> Apps
style Conversion fill:#999999,stroke:#CC0000,color:#000000
style Data fill:#999999,stroke:#CC0000,color:#000000
style Analysis fill:#999999,stroke:#CC0000,color:#000000
style Viz fill:#999999,stroke:#CC0000,color:#000000
style Apps fill:#999999,stroke:#CC0000,color:#000000See the Architecture vignette for detailed diagrams of the data pipeline, function hierarchy, and user journey.
Installation
R-Universe (Recommended)
# Install from r-universe (pre-built binaries, fast)
install.packages("micromort", repos = "https://johngavin.r-universe.dev")GitHub
# Install from GitHub (source)
# install.packages("devtools")
devtools::install_github("JohnGavin/micromort")Concepts
Micromort (Acute Risk)
A micromort is a unit of mortality risk equal to a one-in-a-million probability of death per specific event.
- Unit: 1 micromort = 1/1,000,000 = 0.0001% death probability
- Scope: Per discrete event (e.g., one skydive, one surgery, one flight)
- Sign: Always non-negative (it’s a probability)
- Example: Skydiving has ~8 micromorts per jump (8-in-a-million death chance per jump)
Microlife (Chronic Risk)
A microlife is a unit of life expectancy change equal to 30 minutes of expected lifespan, measured per day of exposure.
- Unit: 1 microlife = 30 minutes of life expectancy
- Scope: Per day of maintaining a habit or exposure
- Sign: Positive (life gained) or negative (life lost)
- Example: Smoking 2 cigarettes daily costs -1 microlife/day (losing 30 mins of life expectancy each day you smoke)
Conversion
1 micromort ≈ 0.7 microlives (assuming 40 years remaining life expectancy). The conversion scales linearly with remaining life expectancy:
| Remaining life expectancy | 1 micromort ≈ |
|---|---|
| 10 years | 0.18 microlives |
| 20 years | 0.35 microlives |
| 40 years (default) | 0.70 microlives |
| 60 years | 1.05 microlives |
Use lle(prob, life_expectancy = ...) and as_microlife() to convert at any age. See the Age-Based Hazard Rates section for daily micromort exposure by age.
Morbidity Metrics
Micromorts and microlives focus on mortality. For quality-of-life impacts:
- QALY (Quality-Adjusted Life Year): 1 year of perfect health. Used in healthcare economics.
-
DALY (Disability-Adjusted Life Years): Disease burden combining:
- YLL (Years of Life Lost): Premature mortality component
- YLD (Years Lived with Disability): Morbidity component
- QALD (Quality-Adjusted Life Days): 1 day of perfect health. For common illnesses.
See the Introduction vignette for detailed examples and the Glossary for all acronym definitions.
Quick Start
Load the Datasets
library(micromort)
# Load acute risks (micromorts per event)
acute <- load_acute_risks()
nrow(acute)
#> [1] 62
# Load chronic risks (microlives per day)
chronic <- load_chronic_risks()
nrow(chronic)
#> [1] 22
# Load source registry
sources <- load_sources()
nrow(sources)
#> [1] 14Acute Risks (Micromorts per Event)
# Convert a probability to micromorts
# Example: 1 in 10,000 chance = 100 micromorts
as_micromort(1/10000)
#> [1] 100
# Top 10 riskiest activities (micromorts per event/period)
acute |>
dplyr::select(activity, micromorts, category, period) |>
head(10)
#> # A tibble: 10 × 4
#> activity micromorts category period
#> <chr> <dbl> <chr> <chr>
#> 1 Mt. Everest ascent 37932 Mountaineering per ascent
#> 2 Himalayan mountaineering 12000 Mountaineering per expedi…
#> 3 COVID-19 infection (unvaccinated) 10000 Disease per infect…
#> 4 Spanish flu infection 3000 Disease per infect…
#> 5 Matterhorn ascent 2840 Mountaineering per ascent
#> 6 Living in US during COVID-19 (Jul 2020) 500 Disease per month
#> 7 Living (one day, age 90) 463 Daily Life per day
#> 8 Base jumping (per jump) 430 Sport per event
#> 9 First day of life (newborn) 430 Daily Life per day
#> 10 COVID-19 unvaccinated (age 80+) 234 COVID-19 11 weeks (…Caption: Micromorts represent probability of death per event. Higher values = higher risk per occurrence.
Chronic Risks (Microlives per Day)
# Factors that reduce life expectancy (microlives lost per day)
chronic |>
dplyr::filter(direction == "loss") |>
dplyr::select(factor, microlives_per_day, category) |>
head(10)
#> # A tibble: 10 × 3
#> factor microlives_per_day category
#> <chr> <dbl> <chr>
#> 1 Smoking 20 cigarettes -10 Smoking
#> 2 Smoking 10 cigarettes -5 Smoking
#> 3 Being male (vs female) -4 Demographics
#> 4 Being 15 kg overweight -3 Weight
#> 5 Being 10 kg overweight -2 Weight
#> 6 4th-5th alcoholic drink -2 Alcohol
#> 7 Smoking 2 cigarettes -1 Smoking
#> 8 Being 5 kg overweight -1 Weight
#> 9 2nd-3rd alcoholic drink -1 Alcohol
#> 10 Red meat (1 portion/day) -1 DietCaption: Microlives per day. Negative values = life expectancy loss; positive = gain. Effects accumulate daily.
Visualize Risks
# Filter to show only activities with micromorts >= 1 for clarity on log scale
plot_risks(common_risks() |> dplyr::filter(micromorts >= 1))
#> Warning in ggplot2::scale_y_log10(labels = scales::comma, limits = c(0.01, : log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.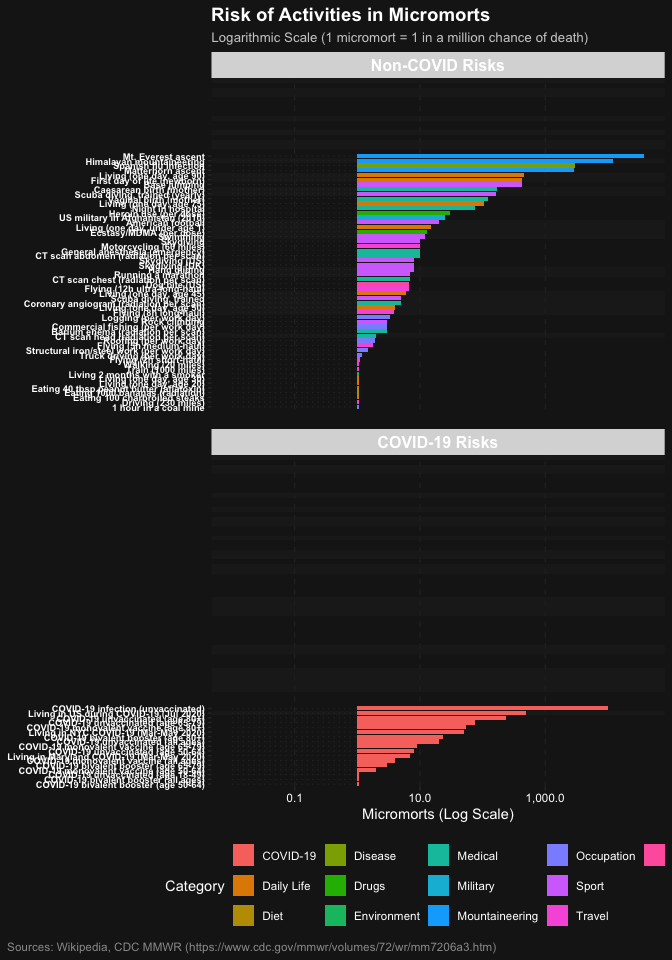
Analysis Functions
Compare Lifestyle Interventions
# Compare quitting smoking vs losing weight (microlives gained per day)
compare_interventions(list(
"Quit 10 cigarettes/day" = list(factor = "Smoking 10 cigarettes", change = -1),
"Lose 5kg" = list(factor = "Being 5 kg overweight", change = -1)
))
#> # A tibble: 2 × 7
#> intervention factor original_ml_per_day change net_ml_per_day annual_days
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 Quit 10 cigarett… Smoki… -5 -1 -5 -38
#> 2 Lose 5kg Being… -1 -1 -1 -7.6
#> # ℹ 1 more variable: lifetime_years <dbl>Calculate Baseline Risk by Age
# Daily baseline mortality risk at age 35 (micromorts per day just from being alive)
daily_hazard_rate(35)
#> # A tibble: 1 × 6
#> age sex daily_prob micromorts microlives_consumed interpretation
#> <dbl> <chr> <dbl> <dbl> <dbl> <chr>
#> 1 35 male 0.00000296 3 0.05 At age 35 (male): 3.0 m…Lifestyle Tradeoffs
# How much exercise offsets smoking? (in microlives per day)
lifestyle_tradeoff("Smoking 2 cigarettes", "20 min moderate exercise")
#> # A tibble: 1 × 6
#> bad_habit bad_ml_per_day good_habit good_ml_per_day units_needed
#> <chr> <dbl> <chr> <dbl> <dbl>
#> 1 Smoking 2 cigarettes -1 20 min moder… 2 0.5
#> # ℹ 1 more variable: interpretation <chr>API Access
Launch the REST API for programmatic access:
launch_api()
# API available at http://localhost:8080
# Swagger docs at http://localhost:8080/__docs__/Core endpoints (30 total — see REST API vignette for full reference):
-
GET /v1/risks/acute— Acute risks (micromorts per event) -
GET /v1/risks/chronic— Chronic risks (microlives per day) -
GET /v1/risks/cancer— Cancer mortality by type/sex/age -
GET /v1/analysis/equivalence— Risk equivalence lookup -
GET /v1/convert/hazard-rate?age=35— Daily hazard rate -
GET /v1/sources— Source registry -
GET /health— Health check
Risk Quiz
Play in browser — runs via WebR/Shinylive (30-60s initial load). Or locally:
micromort::launch_quiz()Data Sources
| Source | Type | Data |
|---|---|---|
| Wikipedia: Micromort | Encyclopedia | ~50 acute risks |
| Wikipedia: Microlife | Encyclopedia | ~20 chronic risks |
| micromorts.rip | Database | ~45 acute risks |
| CDC MMWR | Government | COVID vaccine data |
| Spiegelhalter (2012) BMJ | Academic | Microlife framework |
| SEER Cancer Statistics | Government | Cancer mortality by type/sex |
Project Structure
Click to expand project tree
#> .
#> ├── DESCRIPTION
#> ├── LICENSE
#> ├── LICENSE.md
#> ├── NAMESPACE
#> ├── R
#> │ ├── activity_descriptions.R
#> │ ├── api.R
#> │ ├── atomic_risks.R
#> │ ├── dashboard.R
#> │ ├── data.R
#> │ ├── dev
#> │ │ ├── issues
#> │ │ └── verify_pkgdown_urls.R
#> │ ├── diagrams.R
#> │ ├── micromort.R
#> │ ├── models.R
#> │ ├── quiz.R
#> │ ├── radiation_profiles.R
#> │ ├── regional.R
#> │ ├── risk_equivalence.R
#> │ ├── risks.R
#> │ ├── tar_plans
#> │ │ ├── plan_data_acquisition.R
#> │ │ ├── plan_documentation.R
#> │ │ ├── plan_export.R
#> │ │ ├── plan_logging.R
#> │ │ ├── plan_normalization.R
#> │ │ ├── plan_qa_gates.R
#> │ │ ├── plan_telemetry.R
#> │ │ ├── plan_validation.R
#> │ │ └── plan_vignette_outputs.R
#> │ └── visualization.R
#> ├── README.md
#> ├── README.qmd
#> ├── README.rmarkdown
#> ├── README_files
#> │ └── libs
#> │ ├── bootstrap
#> │ ├── clipboard
#> │ └── quarto-html
#> ├── box
#> │ ├── api
#> │ │ ├── __init__.R
#> │ │ └── endpoints.R
#> │ ├── dashboard
#> │ │ ├── __init__.R
#> │ │ ├── server.R
#> │ │ └── ui.R
#> │ ├── data
#> │ │ ├── __init__.R
#> │ │ ├── loaders.R
#> │ │ ├── parsers.R
#> │ │ └── schemas.R
#> │ └── models
#> │ ├── __init__.R
#> │ ├── compare.R
#> │ └── hazard.R
#> ├── check
#> │ ├── micromort.Rcheck
#> │ │ ├── 00_pkg_src
#> │ │ ├── 00check.log
#> │ │ ├── 00install.out
#> │ │ ├── R_check_bin
#> │ │ ├── micromort
#> │ │ ├── micromort-Ex.R
#> │ │ ├── micromort-Ex.Rout
#> │ │ ├── micromort-Ex.pdf
#> │ │ └── micromort-Ex.timings
#> │ └── micromort_0.1.0.tar.gz
#> ├── data-raw
#> │ ├── 01_extract_current_data.R
#> │ ├── 02_regional_life_expectancy.R
#> │ ├── 02_regional_life_expectancy_sample.R
#> │ ├── 03_osha_occupational_risks.R
#> │ ├── 04_road_traffic_mortality.R
#> │ ├── 05_global_homicide_rates.R
#> │ ├── README_regional_data.md
#> │ ├── generate_og_image.R
#> │ ├── generate_quiz_csv.R
#> │ └── sources
#> │ ├── acute_risks_base.csv
#> │ ├── chronic_risks_base.csv
#> │ ├── covid_vaccine_rr.csv
#> │ ├── demographic_factors.csv
#> │ └── risk_sources.csv
#> ├── default.R
#> ├── default.nix
#> ├── default.sh
#> ├── docs
#> │ ├── 404.html
#> │ ├── 404.md
#> │ ├── LICENSE-text.html
#> │ ├── LICENSE-text.md
#> │ ├── LICENSE.html
#> │ ├── LICENSE.md
#> │ ├── articles
#> │ │ ├── architecture.html
#> │ │ ├── architecture.md
#> │ │ ├── architecture_files
#> │ │ ├── confounding.html
#> │ │ ├── confounding.md
#> │ │ ├── confounding_files
#> │ │ ├── data_reliability.html
#> │ │ ├── data_reliability.md
#> │ │ ├── data_reliability_files
#> │ │ ├── index.html
#> │ │ ├── index.md
#> │ │ ├── introduction.html
#> │ │ ├── introduction.md
#> │ │ ├── introduction_files
#> │ │ ├── palatable_units.html
#> │ │ ├── palatable_units.md
#> │ │ ├── palatable_units_files
#> │ │ ├── quiz_shinylive.html
#> │ │ ├── quiz_shinylive_files
#> │ │ ├── regional_variation.html
#> │ │ ├── regional_variation.md
#> │ │ ├── regional_variation_files
#> │ │ ├── rest_api.html
#> │ │ ├── rest_api.md
#> │ │ ├── rest_api_files
#> │ │ ├── risk_equivalence.html
#> │ │ ├── risk_equivalence.md
#> │ │ ├── risk_equivalence_files
#> │ │ ├── shinylive-sw.js
#> │ │ ├── telemetry.html
#> │ │ ├── telemetry.md
#> │ │ └── telemetry_files
#> │ ├── authors.html
#> │ ├── authors.md
#> │ ├── deps
#> │ │ ├── bootstrap-5.3.1
#> │ │ ├── bootstrap-toc-1.0.1
#> │ │ ├── clipboard.js-2.0.11
#> │ │ ├── data-deps.txt
#> │ │ ├── font-awesome-6.5.2
#> │ │ ├── headroom-0.11.0
#> │ │ ├── jquery-3.6.0
#> │ │ └── search-1.0.0
#> │ ├── extra.css
#> │ ├── extra.js
#> │ ├── index.html
#> │ ├── index.md
#> │ ├── katex-auto.js
#> │ ├── lightswitch.js
#> │ ├── link.svg
#> │ ├── llms.txt
#> │ ├── news
#> │ ├── pkgdown.js
#> │ ├── pkgdown.yml
#> │ ├── reference
#> │ │ ├── activity_descriptions.html
#> │ │ ├── activity_descriptions.md
#> │ │ ├── acute_risks.html
#> │ │ ├── acute_risks.md
#> │ │ ├── annual_risk_budget.html
#> │ │ ├── annual_risk_budget.md
#> │ │ ├── as_microlife.html
#> │ │ ├── as_microlife.md
#> │ │ ├── as_micromort.html
#> │ │ ├── as_micromort.md
#> │ │ ├── as_probability.html
#> │ │ ├── as_probability.md
#> │ │ ├── atomic_risks.html
#> │ │ ├── atomic_risks.md
#> │ │ ├── cancer_risks.html
#> │ │ ├── cancer_risks.md
#> │ │ ├── chronic_risks.html
#> │ │ ├── chronic_risks.md
#> │ │ ├── common_risks.html
#> │ │ ├── common_risks.md
#> │ │ ├── compare_interventions.html
#> │ │ ├── compare_interventions.md
#> │ │ ├── conditional_risk.html
#> │ │ ├── conditional_risk.md
#> │ │ ├── covid_vaccine_rr.html
#> │ │ ├── covid_vaccine_rr.md
#> │ │ ├── daily_hazard_rate.html
#> │ │ ├── daily_hazard_rate.md
#> │ │ ├── demographic_factors.html
#> │ │ ├── demographic_factors.md
#> │ │ ├── figures
#> │ │ ├── format_activity_name.html
#> │ │ ├── format_activity_name.md
#> │ │ ├── hedged_portfolio.html
#> │ │ ├── hedged_portfolio.md
#> │ │ ├── index.html
#> │ │ ├── index.md
#> │ │ ├── laggard_regions.html
#> │ │ ├── laggard_regions.md
#> │ │ ├── launch_api.html
#> │ │ ├── launch_api.md
#> │ │ ├── launch_dashboard.html
#> │ │ ├── launch_dashboard.md
#> │ │ ├── launch_quiz.html
#> │ │ ├── launch_quiz.md
#> │ │ ├── libs
#> │ │ ├── lifestyle_tradeoff.html
#> │ │ ├── lifestyle_tradeoff.md
#> │ │ ├── lle.html
#> │ │ ├── lle.md
#> │ │ ├── load_acute_risks.html
#> │ │ ├── load_acute_risks.md
#> │ │ ├── load_chronic_risks.html
#> │ │ ├── load_chronic_risks.md
#> │ │ ├── load_sources.html
#> │ │ ├── load_sources.md
#> │ │ ├── patient_radiation_comparison.html
#> │ │ ├── patient_radiation_comparison.md
#> │ │ ├── plot_risk_components-1.png
#> │ │ ├── plot_risk_components.html
#> │ │ ├── plot_risk_components.md
#> │ │ ├── plot_risks-1.png
#> │ │ ├── plot_risks-2.png
#> │ │ ├── plot_risks-3.png
#> │ │ ├── plot_risks-4.png
#> │ │ ├── plot_risks-5.png
#> │ │ ├── plot_risks.html
#> │ │ ├── plot_risks.md
#> │ │ ├── plot_risks_interactive.html
#> │ │ ├── plot_risks_interactive.md
#> │ │ ├── prepare_risks_plot-1.png
#> │ │ ├── prepare_risks_plot.html
#> │ │ ├── prepare_risks_plot.md
#> │ │ ├── quiz_pairs.html
#> │ │ ├── quiz_pairs.md
#> │ │ ├── radiation_profiles.html
#> │ │ ├── radiation_profiles.md
#> │ │ ├── regional_life_expectancy.html
#> │ │ ├── regional_life_expectancy.md
#> │ │ ├── regional_mortality_multiplier.html
#> │ │ ├── regional_mortality_multiplier.md
#> │ │ ├── risk_components.html
#> │ │ ├── risk_components.md
#> │ │ ├── risk_data_sources.html
#> │ │ ├── risk_data_sources.md
#> │ │ ├── risk_equivalence.html
#> │ │ ├── risk_equivalence.md
#> │ │ ├── risk_exchange_matrix.html
#> │ │ ├── risk_exchange_matrix.md
#> │ │ ├── risk_for_duration.html
#> │ │ ├── risk_for_duration.md
#> │ │ ├── risk_sources.html
#> │ │ ├── risk_sources.md
#> │ │ ├── theme_micromort_dark-1.png
#> │ │ ├── theme_micromort_dark.html
#> │ │ ├── theme_micromort_dark.md
#> │ │ ├── vaccination_risks.html
#> │ │ ├── vaccination_risks.md
#> │ │ ├── value_of_micromort.html
#> │ │ ├── value_of_micromort.md
#> │ │ ├── vanguard_regions.html
#> │ │ └── vanguard_regions.md
#> │ ├── search.json
#> │ ├── sitemap.xml
#> │ └── tutorials
#> ├── inst
#> │ ├── dashboard
#> │ │ └── about.md
#> │ ├── extdata
#> │ │ ├── acute_risks.parquet
#> │ │ ├── chronic_risks.parquet
#> │ │ ├── logs
#> │ │ ├── regional_life_expectancy.parquet
#> │ │ ├── risk_sources.parquet
#> │ │ └── vignettes
#> │ └── plumber
#> │ └── api.R
#> ├── man
#> │ ├── activity_descriptions.Rd
#> │ ├── acute_risks.Rd
#> │ ├── annual_risk_budget.Rd
#> │ ├── as_microlife.Rd
#> │ ├── as_micromort.Rd
#> │ ├── as_probability.Rd
#> │ ├── atomic_risks.Rd
#> │ ├── cancer_risks.Rd
#> │ ├── chronic_risks.Rd
#> │ ├── common_risks.Rd
#> │ ├── compare_interventions.Rd
#> │ ├── conditional_risk.Rd
#> │ ├── covid_vaccine_rr.Rd
#> │ ├── daily_hazard_rate.Rd
#> │ ├── demographic_factors.Rd
#> │ ├── figures
#> │ │ ├── README-plot-1.png
#> │ │ ├── logo-candidates
#> │ │ ├── logo.png
#> │ │ └── og-image.png
#> │ ├── format_activity_name.Rd
#> │ ├── hedged_portfolio.Rd
#> │ ├── laggard_regions.Rd
#> │ ├── launch_api.Rd
#> │ ├── launch_dashboard.Rd
#> │ ├── launch_quiz.Rd
#> │ ├── lifestyle_tradeoff.Rd
#> │ ├── lle.Rd
#> │ ├── load_acute_risks.Rd
#> │ ├── load_chronic_risks.Rd
#> │ ├── load_sources.Rd
#> │ ├── patient_radiation_comparison.Rd
#> │ ├── plot_risk_components.Rd
#> │ ├── plot_risks.Rd
#> │ ├── plot_risks_interactive.Rd
#> │ ├── prepare_risks_plot.Rd
#> │ ├── quiz_pairs.Rd
#> │ ├── radiation_profiles.Rd
#> │ ├── regional_life_expectancy.Rd
#> │ ├── regional_mortality_multiplier.Rd
#> │ ├── risk_components.Rd
#> │ ├── risk_data_sources.Rd
#> │ ├── risk_equivalence.Rd
#> │ ├── risk_exchange_matrix.Rd
#> │ ├── risk_for_duration.Rd
#> │ ├── risk_sources.Rd
#> │ ├── theme_micromort_dark.Rd
#> │ ├── vaccination_risks.Rd
#> │ ├── value_of_micromort.Rd
#> │ └── vanguard_regions.Rd
#> ├── nix-shell-root
#> ├── package.nix
#> ├── pkgdown
#> │ ├── extra.css
#> │ └── extra.js
#> ├── plans
#> │ ├── PLAN_consistency_refactor.md
#> │ ├── PLAN_regional_longevity.md
#> │ ├── PLAN_risk_equivalence_dashboard.md
#> │ └── PLAN_vignette_targets_refactor.md
#> ├── push_to_cachix.sh
#> ├── tests
#> │ └── testthat
#> │ ├── test-adversarial.R
#> │ ├── test-api.R
#> │ ├── test-atomic-risks.R
#> │ ├── test-diagrams.R
#> │ ├── test-quiz.R
#> │ ├── test-radiation-profiles.R
#> │ ├── test-risk-components.R
#> │ ├── test-risk-equivalence.R
#> │ └── test-visualization.R
#> └── vignettes
#> ├── _extensions
#> │ └── quarto-ext
#> ├── _quarto.yml
#> ├── architecture.qmd
#> ├── architecture_files
#> ├── confounding.qmd
#> ├── data_reliability.qmd
#> ├── introduction.qmd
#> ├── palatable_units.qmd
#> ├── quiz_shinylive.qmd
#> ├── quiz_shinylive_files
#> │ └── libs
#> ├── regional_variation.qmd
#> ├── rest_api.qmd
#> ├── risk_equivalence.qmd
#> ├── risk_equivalence_files
#> ├── shinylive-sw.js
#> └── telemetry.qmdContributing
Contributions are welcome! Please:
- Report issues at GitHub Issues
- Submit PRs following the tidyverse style guide
-
Add data - New risk sources welcome! Include:
- Source URL with citation
- Units (micromorts per event OR microlives per day)
- Period specification (per jump, per day, per year, etc.)
Development Setup
Requires Nix (the ./default.sh script builds a reproducible R environment via Nix):
Glossary
| Acronym | Full Name | Definition |
|---|---|---|
| DALY | Disability-Adjusted Life Year | Disease burden = YLL + YLD. 1 DALY = 1 year of healthy life lost. |
| LLE | Loss of Life Expectancy | Expected lifespan reduction from a risk, in minutes. See lle(). |
| QALD | Quality-Adjusted Life Day | 1 day of perfect health. Useful for short-duration conditions. |
| QALY | Quality-Adjusted Life Year | 1 year of perfect health. Used in healthcare cost-effectiveness. |
| VSL | Value of Statistical Life | Monetary value society places on preventing one death (~$10M USD). See value_of_micromort(). |
| YLD | Years Lived with Disability | Morbidity component of DALY. Time spent in impaired health. |
| YLL | Years of Life Lost | Mortality component of DALY. Premature death relative to standard life expectancy. |
References
- Howard RA (1980). “On Making Life and Death Decisions.” Societal Risk Assessment.
- Spiegelhalter D (2012). “Using speed of ageing and ‘microlives’.” BMJ 345:e8223. DOI: 10.1136/bmj.e8223
- Blastland M, Spiegelhalter D (2013). The Norm Chronicles: Stories and Numbers About Danger.
