|
api_pathways
|
|
|
|
code_dd_explore_dict
|
|
|
|
code_eda_aggregation
|
|
|
|
api_clinical
|
|
|
|
api_cytogenetics
|
|
|
|
api_data_dict
|
|
|
|
api_de_results
|
|
|
|
api_index
|
|
|
|
api_survival
|
|
|
|
code_api_python
|
|
|
|
code_readme_quick_start
|
|
|
|
readme_project_tree
|
|
|
|
vig_api_endpoints_dt
|
|
|
|
code_api_curl
|
|
|
|
code_eda_simple_query
|
|
|
|
shiny_survival_index
|
|
|
|
code_api_r
|
|
|
|
code_dd_load_data
|
|
|
|
pkgctx_self
|
|
|
|
code_parsed_dd_explore_dict
|
|
|
|
code_parsed_eda_aggregation
|
|
|
|
save_api_files
|
|
|
|
code_parsed_api_python
|
|
|
|
code_parsed_readme_quick_start
|
|
|
|
code_parsed_api_curl
|
|
|
|
code_parsed_eda_simple_query
|
|
|
|
code_parsed_api_r
|
|
|
|
code_parsed_dd_load_data
|
|
|
|
doc_examples_validation
|
|
|
|
pkgctx_deps
|
|
error in running command. ! pkgctx failed for dplyr ℹ Exit code: 127. error in running command. ! pkgctx failed for targets ℹ Exit code: 127. error in running command. ! pkgctx failed for DBI ℹ Exit code: 127. error in running command. ! pkgctx failed for duckdb ℹ Exit code: 127. error in running command. ! pkgctx failed for arrow ℹ Exit code: 127. error in running command. ! pkgctx failed for cli ℹ Exit code: 127
|
|
cyto_viz_data
|
|
|
|
cyto_cooccurrence
|
|
|
|
vig_cyto_oncoprint
|
|
|
|
vig_cyto_cooccurrence_heatmap
|
|
|
|
cyto_frequency_summary
|
|
|
|
clinical_data
|
|
|
|
eda_missing_summary
|
|
|
|
eda_iss_summary
|
|
NAs introduced by coercion
|
|
eda_biospecimen_summary
|
|
|
|
vig_biospecimen_overview_text
|
|
|
|
eda_integration_summary
|
|
|
|
vig_integration_text
|
|
|
|
vig_missing_data_text
|
|
|
|
eda_duckdb_demo
|
|
|
|
vig_missing_data_table
|
|
|
|
shiny_km_os_by_response
|
|
|
|
km_by_markers
|
|
|
|
shiny_cox_os_age_stage
|
|
|
|
shiny_risk_os_by_response
|
|
|
|
vig_km_markers
|
|
|
|
km_by_iss
|
|
|
|
shiny_pfs_data
|
|
|
|
cox_basic
|
|
|
|
shiny_risk_os_by_iss
|
|
|
|
shiny_km_pfs_by_iss
|
|
|
|
shiny_cox_os_age_risk
|
|
|
|
shiny_risk_pfs_by_iss
|
|
|
|
km_by_risk
|
|
|
|
km_overall
|
|
|
|
shiny_cox_os_full
|
|
|
|
vig_km_risk
|
|
|
|
cox_full
|
|
|
|
vig_km_overall
|
|
|
|
vig_km_iss
|
|
|
|
shiny_km_pfs_by_risk
|
|
|
|
shiny_km_pfs_overall
|
|
|
|
shiny_risk_os_by_risk
|
|
|
|
shiny_risk_pfs_by_risk
|
|
|
|
shiny_risk_pfs_overall
|
|
|
|
shiny_risk_os_overall
|
|
|
|
vig_forest_plot
|
|
|
|
raw_rnaseq
|
|
|
|
vig_dd_rnaseq_biotype_table
|
|
|
|
vig_dd_rnaseq_text
|
|
|
|
eda_rnaseq_summary
|
|
|
|
expression_data_clean
|
|
|
|
qc_metrics
|
|
|
|
eda_qc_summary
|
|
|
|
vig_qc_text
|
|
|
|
vig_rnaseq_summary_text
|
|
|
|
vig_rnaseq_quality_text
|
|
|
|
vig_qc_table
|
|
|
|
integrated_data
|
|
No matching sample IDs found between clinical and expression data
|
|
data_quality_report
|
|
|
|
filtered_data
|
|
|
|
vig_dd_biospecimen_text
|
|
|
|
vig_dd_clinical_text
|
|
|
|
vig_dd_clinical_table
|
|
|
|
vig_clinical_column_info
|
|
|
|
vig_rnaseq_sample_stats
|
|
|
|
vig_filtered_summary
|
|
|
|
normalized_data
|
|
|
|
vst_counts
|
|
|
|
pca_data
|
|
|
|
vig_pca_plot
|
|
|
|
gsea_hallmark
|
|
|
|
enrichment_dotplot_data
|
|
|
|
enrichment_barplot_data
|
|
|
|
vig_gsea_dotplot
|
|
|
|
vig_ora_barplot
|
|
|
|
vig_paired_de_text
|
|
|
|
vig_paired_de_plot
|
|
|
|
dag_layer_validation
|
|
|
|
code_vig_annotated_de_table
|
|
|
|
config
|
|
|
|
code_vig_cyto_frequency_table
|
|
|
|
vig_config_text
|
|
|
|
code_vig_cox_comparison
|
|
|
|
code_vig_git_info
|
|
|
|
code_vig_duckdb_demo_agg
|
|
|
|
code_vig_cox_basic_table
|
|
|
|
code_vig_duckdb_demo_simple
|
|
|
|
code_vig_iss_age_table
|
|
|
|
code_vig_ph_test_table
|
|
|
|
code_vig_de_method_table
|
|
|
|
code_vig_cyto_cooccurrence_table
|
|
|
|
code_vig_count_distribution_plot
|
|
|
|
code_vig_gender_table
|
|
|
|
code_vig_iss_table
|
|
|
|
vig_git_info
|
|
|
|
code_vig_vital_table
|
|
|
|
code_vig_race_table
|
|
|
|
code_vig_cox_full_table
|
|
|
|
vig_units_table
|
|
|
|
glossary_table
|
|
|
|
vig_dd_column_mappings
|
|
|
|
vig_dd_dictionary_table
|
|
|
|
vig_msigdb_stats
|
|
|
|
vig_pipeline_dag
|
|
|
|
cytogenetic_data
|
|
|
|
clinical_data_clean
|
|
NAs introduced by coercion
|
|
survival_data
|
|
NAs introduced by coercion
|
|
vig_dd_biospecimen_table
|
|
|
|
vig_dd_rnaseq_sample_table
|
|
|
|
code_vig_age_histogram
|
|
|
|
code_vig_de_method_barplot
|
|
|
|
nix_desc_file
|
|
|
|
nix_desc_deps
|
|
|
|
nix_sync_check
|
|
! DESCRIPTION -> Nix drift detected! ℹ Packages in DESCRIPTION but not in default.nix: TCGAbiolinks, S4Vectors, SummarizedExperiment, edgeR, logger, dplyr, aws.s3, arrow, DBI, duckdb, dbplyr, cli, jsonlite, glue, stringr, tibble, anndataR, AnnotationDbi, …, visNetwork, and here ℹ Run Rscript default.R to regenerate.
|
|
code_vig_iss_barplot
|
|
|
|
vig_genes_detected_hist
|
|
|
|
treatment_data_clean
|
|
|
|
vig_library_size_boxplot
|
|
|
|
treatment_summary
|
|
|
|
vig_iss_age_table
|
|
|
|
vig_iss_distribution_text
|
|
|
|
vig_iss_table
|
|
|
|
vig_duckdb_demo_agg
|
|
|
|
vig_duckdb_demo_simple
|
|
|
|
vig_iss_barplot
|
|
|
|
vig_cyto_cooccurrence_table
|
|
|
|
vig_samples_per_patient_hist
|
|
|
|
vig_cyto_frequency_table
|
|
|
|
vig_sample_type_barplot
|
|
|
|
vig_gene_correlations
|
|
|
|
eda_pca_summary
|
|
|
|
vig_pca_biplot_vital
|
|
|
|
vig_cox_basic_table
|
|
|
|
vig_cox_comparison
|
|
|
|
vig_ph_test_table
|
|
|
|
vig_cox_full_table
|
|
|
|
de_report
|
|
|
|
summary_report
|
|
|
|
vig_pca_biplot
|
|
|
|
vig_gene_expression_dist
|
|
|
|
vig_gene_correlation_plot
|
|
|
|
eda_clinical_summary
|
|
|
|
skill_source_files
|
|
|
|
skill_drift_check
|
|
! gdc-genomics skill may be stale! ℹ R source files changed since skill was last synced. ℹ Review ~/.claude/skills/gdc-genomics/references/ and update.
|
|
vig_gender_table
|
|
|
|
vig_age_histogram
|
|
|
|
vig_survival_endpoints
|
|
|
|
vig_vital_table
|
|
|
|
vig_race_table
|
|
|
|
vig_clinical_demographics_text
|
|
|
|
vig_gene_expr_subtype
|
|
|
|
limma_results
|
|
|
|
vig_gene_km_expression
|
|
|
|
vig_km_overall_text
|
|
|
|
edger_results
|
|
|
|
deseq2_paired_results
|
|
the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, possibly due to insufficient numeric precision. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, possibly due to insufficient numeric precision. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, unable to sufficiently decrease the function value. the line search routine failed, possibly due to insufficient numeric precision. the line search routine failed, unable to su
|
|
deseq2_results
|
|
|
|
ma_plot_data
|
|
|
|
volcano_plot_data
|
|
|
|
vig_ma_plot
|
|
|
|
gene_annotation
|
|
|
|
gsea_kegg
|
✖ Package msigdbr required but not installed ℹ Run ‘R/dev/create_msigdb_data.R’ to generate local gene sets
|
|
|
de_method_summary
|
|
|
|
vig_de_method_barplot
|
|
|
|
de_results_annotated
|
|
|
|
vig_annotated_de_table
|
|
|
|
consensus_de_genes
|
|
|
|
top_de_heatmap_data
|
|
|
|
vig_volcano_plot
|
|
|
|
vig_de_method_table
|
|
|
|
vig_heatmap_plot
|
|
|
|
vig_consensus_text
|
|
|
|
all_outdated
|
|
|
|
to_build
|
|
|
|
exclude
|
|
|
|
vig_git_changelog_summary
|
|
|
|
vig_changes_by_type
|
|
|
|
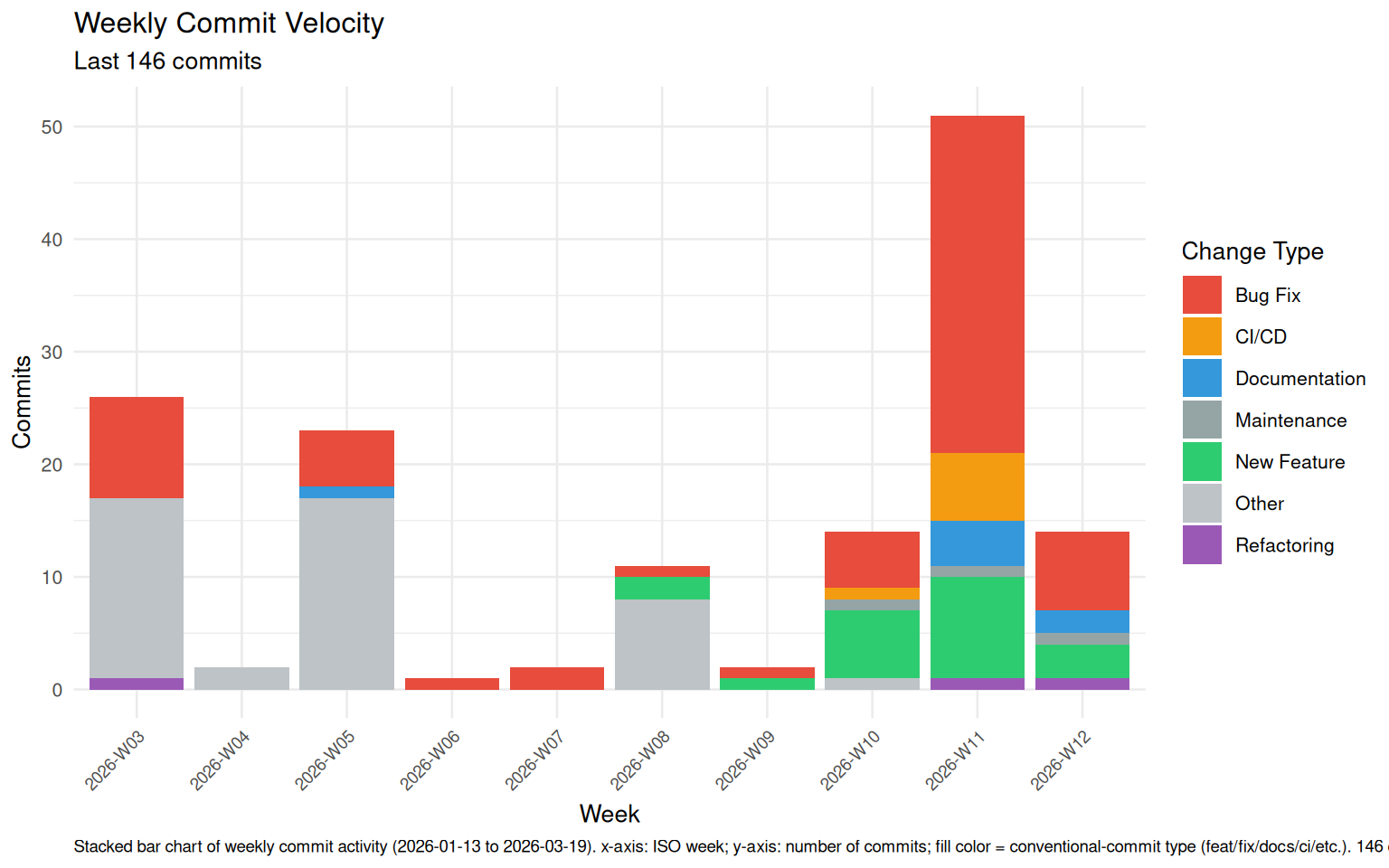
vig_commit_velocity
|
|
|
|
vig_changes_by_file_category
|
|
|
|
gene_correlations
|
|
|
|
km_by_expression
|
|
|
|
vig_km_by_expression
|
|
|
|
expr_by_subtype
|
|
|
|
vig_expr_by_subtype
|
|
|
|
vig_git_changelog
|
|
|
|
vig_build_status
|
|
|
|
vig_outdated_list
|
|
|
|
vig_github_activity
|
|
|
|
vig_github_activity_table
|
|
|
|
ora_hallmark
|
✖ Package msigdbr required but not installed ℹ Run ‘R/dev/create_msigdb_data.R’ to generate local gene sets
|
|
|
vig_pipeline_summary
|
|
|
|
vig_timing_table
|
|
|
|
vig_errors_table
|
|
|
|
vig_codebase_metrics
|
|
|
|
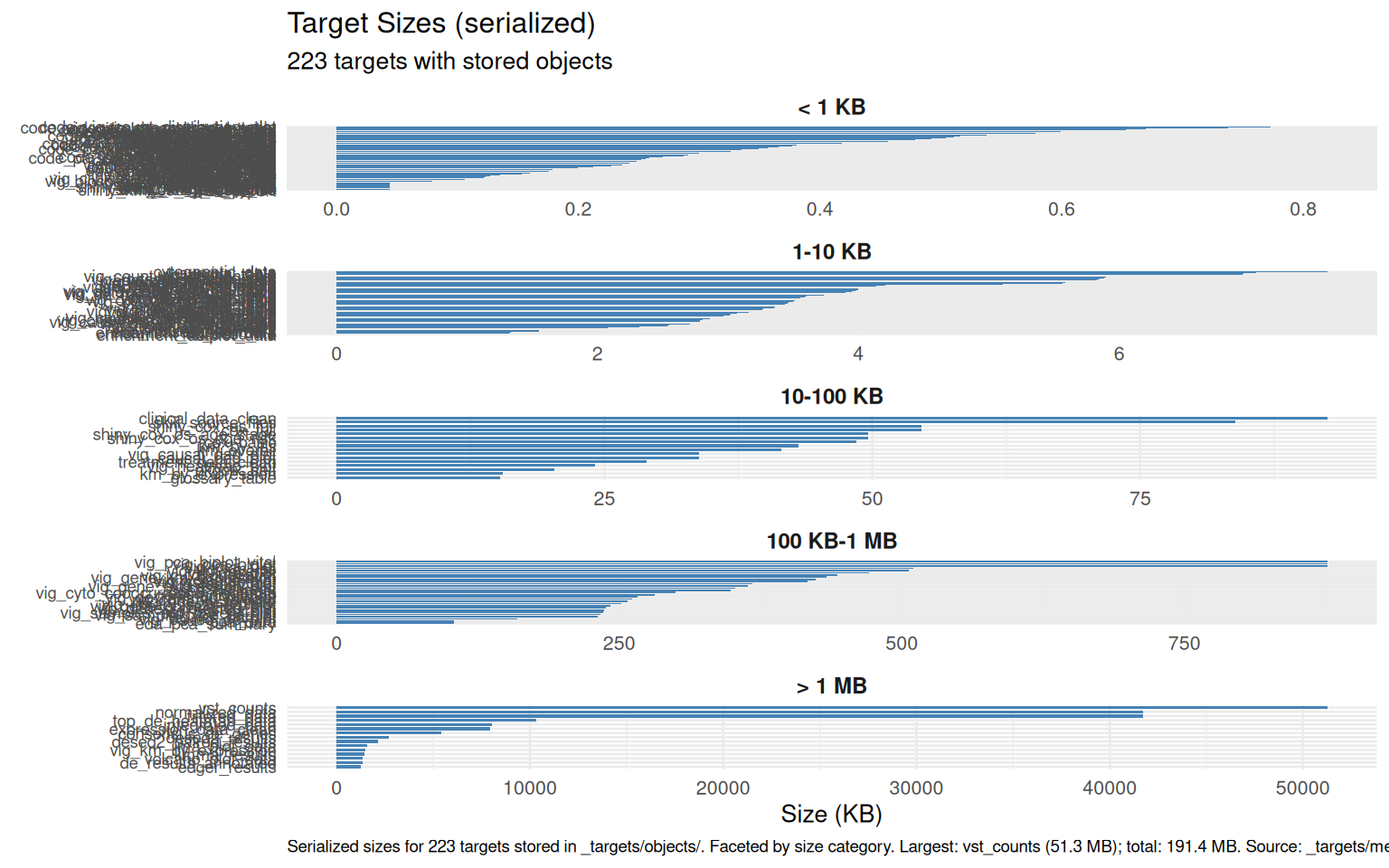
vig_size_plot
|
|
|
|
response_levels
|
|
|
|
get_commpass_data_dictionary
|
|
|
|
setup_logging
|
|
|
|
run_pkgctx
|
|
|
|
make_iss_table
|
|
|
|
generate_api_index
|
|
|
|
download_aws_data
|
|
|
|
download_clinical_data
|
|
|
|
parse_git_changelog
|
|
|
|
make_iss_barplot
|
|
|
|
export_h5ad
|
|
|
|
make_ph_test_table
|
|
|
|
update.packages
|
|
|
|
get_counts_assay
|
|
|
|
make_cyto_frequency_table
|
|
|
|
find_consensus_genes
|
|
|
|
find_expr_col
|
|
|
|
generate_api_endpoint
|
|
|
|
placeholder_results
|
|
|
|
save_timestamped
|
|
|
|
plot_enrichment_dotplot
|
|
|
|
compute_pca
|
|
|
|
annotate_via_orgdb
|
|
|
|
resolve_parquet_path
|
|
|
|
prepare_survival_data
|
|
|
|
plot_pca
|
|
|
|
api_serve
|
|
|
|
make_age_histogram
|
|
|
|
find_lfc_col
|
|
|
|
find_padj_col
|
|
|
|
prepare_coldata
|
|
|
|
standardize_regimen
|
|
|
|
correlate_genes
|
|
|
|
plot_gene_correlation
|
|
|
|
make_duckdb_demo_agg
|
|
|
|
parse_code_example
|
|
|
|
parse_commpass_barcodes
|
|
|
|
api_get_de_results
|
|
|
|
make_annotated_de_table
|
|
|
|
plot_forest
|
|
|
|
resolve_gene_id
|
|
|
|
summarize_treatment
|
|
|
|
plot_expression_by_subtype
|
|
|
|
check_dependencies
|
|
|
|
build_gene_ranks
|
|
|
|
classify_cytogenetic_risk
|
|
|
|
parse_cytogenetic_status
|
|
|
|
summarize_cytogenetics
|
|
|
|
format_file_size
|
|
|
|
show_target
|
|
|
|
make_gender_table
|
|
|
|
load_local_gene_sets
|
|
|
|
download_s3_subset
|
|
|
|
run_kaplan_meier
|
|
|
|
make_race_table
|
|
|
|
install.packages
|
|
|
|
classify_regimen
|
|
|
|
integrate_clinical_expression
|
|
|
|
generate_summary_report
|
|
|
|
make_vital_table
|
|
|
|
remove.packages
|
|
|
|
make_de_method_table
|
|
|
|
render_de_report
|
|
|
|
make_iss_age_table
|
|
|
|
strip_plotly
|
|
|
|
run_cox_regression
|
|
|
|
plot_enrichment_barplot
|
|
|
|
api_get_survival
|
|
|
|
validate_pkgdown_articles
|
|
|
|
plot_cooccurrence_heatmap
|
|
|
|
make_cox_comparison
|
|
|
|
annotate_via_msigdbr
|
|
|
|
make_git_info
|
|
|
|
plot_km
|
|
|
|
make_cyto_cooccurrence_table
|
|
|
|
api_list_datasets
|
|
|
|
create_summary_table
|
|
|
|
make_de_method_barplot
|
|
|
|
gene_report
|
|
|
|
make_duckdb_demo_simple
|
|
|
|
plot_cytogenetic_oncoprint
|
|
|
|
format_with_commas
|
|
|
|
extract_risk_table
|
|
|
|
create_project_dirs
|
|
|
|
get_description_deps
|
|
|
|
list_s3_commpass
|
|
|
|
query_commpass_rna
|
|
|
|
make_cox_full_table
|
|
|
|
api_get_pathways
|
|
|
|
get_commpass_clinical
|
|
|
|
api_get_clinical
|
|
|
|
calculate_cooccurrence
|
|
|
|
compute_riss
|
|
|
|
.__global__
|
|
|
|
make_cox_basic_table
|
|
|
|
clean_clinical_data
|
|
|
|
clean_expression_data
|
|
|
|
annotate_de_results
|
|
|
|
standardize_response
|
|
|
|
get_variable_docs
|
|
|
|
save_rnaseq_parquet
|
|
|
|
make_count_distribution_plot
|
|
|
|
extract_counts
|
|
|
|
run_vst
|
|
|
|
normalize_rnaseq
|
|
|
|
summarize_data
|
|
|
|
calculate_qc_metrics
|
|
|
|
query_commpass_parquet
|
|
|
|
get_commpass_tbl
|
|
|
|
summarize_de_methods
|
|
|
|
plot_ma
|
|
|
|
plot_volcano
|
|
|
|
plot_heatmap_de
|
|
|
|
correlate_genes_batch
|
|
|
|
build_paired_coldata
|
|
|
|
extract_cytogenetic_data
|
|
|
|
get_msigdb_gene_sets
|
|
|
|
run_km_by_markers
|
|
|
|
run_km_by_expression
|
|
|
|
annotate_genes
|
|
|
|
clean_treatment_data
|
|
|
|
download_gdc_rnaseq
|
|
|
|
run_edger
|
|
|
|
run_limma
|
|
|
|
filter_low_quality
|
|
|
|
run_deseq2_paired
|
|
|
|
run_ora
|
|
|
|
run_gsea
|
|
|
|
plot_gsea_running_score
|
|
|
|
example_data
|
|
|
|
acquire_commpass_data
|
|
|
|
run_deseq2
|
|
|
|
run_pathway_analysis
|
|
|
|
target_name
|
|
|
|
plan_file
|
|
|
|
file
|
|
|
|
plan_files
|
|
|
|
vig_count_distribution_plot
|
could not find function ‘get_counts_assay’
|
|
|
response_levels
|
|
|
|
get_commpass_data_dictionary
|
|
|
|
setup_logging
|
|
|
|
run_pkgctx
|
|
|
|
make_iss_table
|
|
|
|
generate_api_index
|
|
|
|
download_aws_data
|
|
|
|
download_clinical_data
|
|
|
|
parse_git_changelog
|
|
|
|
make_iss_barplot
|
|
|
|
export_h5ad
|
|
|
|
make_ph_test_table
|
|
|
|
update.packages
|
|
|
|
get_counts_assay
|
|
|
|
make_cyto_frequency_table
|
|
|
|
find_consensus_genes
|
|
|
|
find_expr_col
|
|
|
|
file
|
|
|
|
generate_api_endpoint
|
|
|
|
placeholder_results
|
|
|
|
save_timestamped
|
|
|
|
plot_enrichment_dotplot
|
|
|
|
compute_pca
|
|
|
|
annotate_via_orgdb
|
|
|
|
resolve_parquet_path
|
|
|
|
prepare_survival_data
|
|
|
|
plot_pca
|
|
|
|
api_serve
|
|
|
|
make_age_histogram
|
|
|
|
find_lfc_col
|
|
|
|
find_padj_col
|
|
|
|
prepare_coldata
|
|
|
|
standardize_regimen
|
|
|
|
correlate_genes
|
|
|
|
plot_gene_correlation
|
|
|
|
make_duckdb_demo_agg
|
|
|
|
parse_code_example
|
|
|
|
parse_commpass_barcodes
|
|
|
|
api_get_de_results
|
|
|
|
make_annotated_de_table
|
|
|
|
plot_forest
|
|
|
|
resolve_gene_id
|
|
|
|
summarize_treatment
|
|
|
|
plot_expression_by_subtype
|
|
|
|
check_dependencies
|
|
|
|
build_gene_ranks
|
|
|
|
classify_cytogenetic_risk
|
|
|
|
parse_cytogenetic_status
|
|
|
|
summarize_cytogenetics
|
|
|
|
format_file_size
|
|
|
|
show_target
|
|
|
|
make_gender_table
|
|
|
|
load_local_gene_sets
|
|
|
|
download_s3_subset
|
|
|
|
run_kaplan_meier
|
|
|
|
make_race_table
|
|
|
|
install.packages
|
|
|
|
classify_regimen
|
|
|
|
integrate_clinical_expression
|
|
|
|
plan_file
|
|
|
|
generate_summary_report
|
|
|
|
make_vital_table
|
|
|
|
remove.packages
|
|
|
|
make_de_method_table
|
|
|
|
render_de_report
|
|
|
|
make_iss_age_table
|
|
|
|
strip_plotly
|
|
|
|
run_cox_regression
|
|
|
|
plot_enrichment_barplot
|
|
|
|
api_get_survival
|
|
|
|
validate_pkgdown_articles
|
|
|
|
plot_cooccurrence_heatmap
|
|
|
|
make_cox_comparison
|
|
|
|
annotate_via_msigdbr
|
|
|
|
make_git_info
|
|
|
|
plot_km
|
|
|
|
make_cyto_cooccurrence_table
|
|
|
|
api_list_datasets
|
|
|
|
create_summary_table
|
|
|
|
make_de_method_barplot
|
|
|
|
gene_report
|
|
|
|
make_duckdb_demo_simple
|
|
|
|
plot_cytogenetic_oncoprint
|
|
|
|
format_with_commas
|
|
|
|
extract_risk_table
|
|
|
|
plan_files
|
|
|
|
create_project_dirs
|
|
|
|
get_description_deps
|
|
|
|
list_s3_commpass
|
|
|
|
query_commpass_rna
|
|
|
|
make_cox_full_table
|
|
|
|
api_get_pathways
|
|
|
|
get_commpass_clinical
|
|
|
|
api_get_clinical
|
|
|
|
calculate_cooccurrence
|
|
|
|
compute_riss
|
|
|
|
.__global__
|
|
|
|
make_cox_basic_table
|
|
|
|
clean_clinical_data
|
|
|
|
clean_expression_data
|
|
|
|
annotate_de_results
|
|
|
|
standardize_response
|
|
|
|
get_variable_docs
|
|
|
|
save_rnaseq_parquet
|
|
|
|
make_count_distribution_plot
|
|
|
|
extract_counts
|
|
|
|
run_vst
|
|
|
|
normalize_rnaseq
|
|
|
|
summarize_data
|
|
|
|
calculate_qc_metrics
|
|
|
|
query_commpass_parquet
|
|
|
|
get_commpass_tbl
|
|
|
|
summarize_de_methods
|
|
|
|
plot_ma
|
|
|
|
plot_volcano
|
|
|
|
plot_heatmap_de
|
|
|
|
correlate_genes_batch
|
|
|
|
build_paired_coldata
|
|
|
|
extract_cytogenetic_data
|
|
|
|
get_msigdb_gene_sets
|
|
|
|
run_km_by_markers
|
|
|
|
run_km_by_expression
|
|
|
|
annotate_genes
|
|
|
|
clean_treatment_data
|
|
|
|
download_gdc_rnaseq
|
|
|
|
run_edger
|
|
|
|
run_limma
|
|
|
|
filter_low_quality
|
|
|
|
run_deseq2_paired
|
|
|
|
run_ora
|
|
|
|
run_gsea
|
|
|
|
plot_gsea_running_score
|
|
|
|
example_data
|
|
|
|
acquire_commpass_data
|
|
|
|
run_deseq2
|
|
|
|
run_pathway_analysis
|
|
|
|
vig_pipeline_summary
|
|
|
|
vig_timing_table
|
|
|