Online documentation
This vignette displays pre-computed results for the default gene (CD70). Run the targets pipeline locally for interactive gene-specific analysis.
CD70 — Single Gene Characterization
This report provides a multi-faceted characterization of CD70 across the CoMMpass cohort: expression distribution, survival stratification, cytogenetic context, and co-expression correlations.
Generated with gene_report("CD70"). See ?gene_report for usage.
Expression Distribution
VST-normalized expression values across all samples, showing the distribution shape and central tendency.
 -1.png){width=768}
-1.png){width=768}
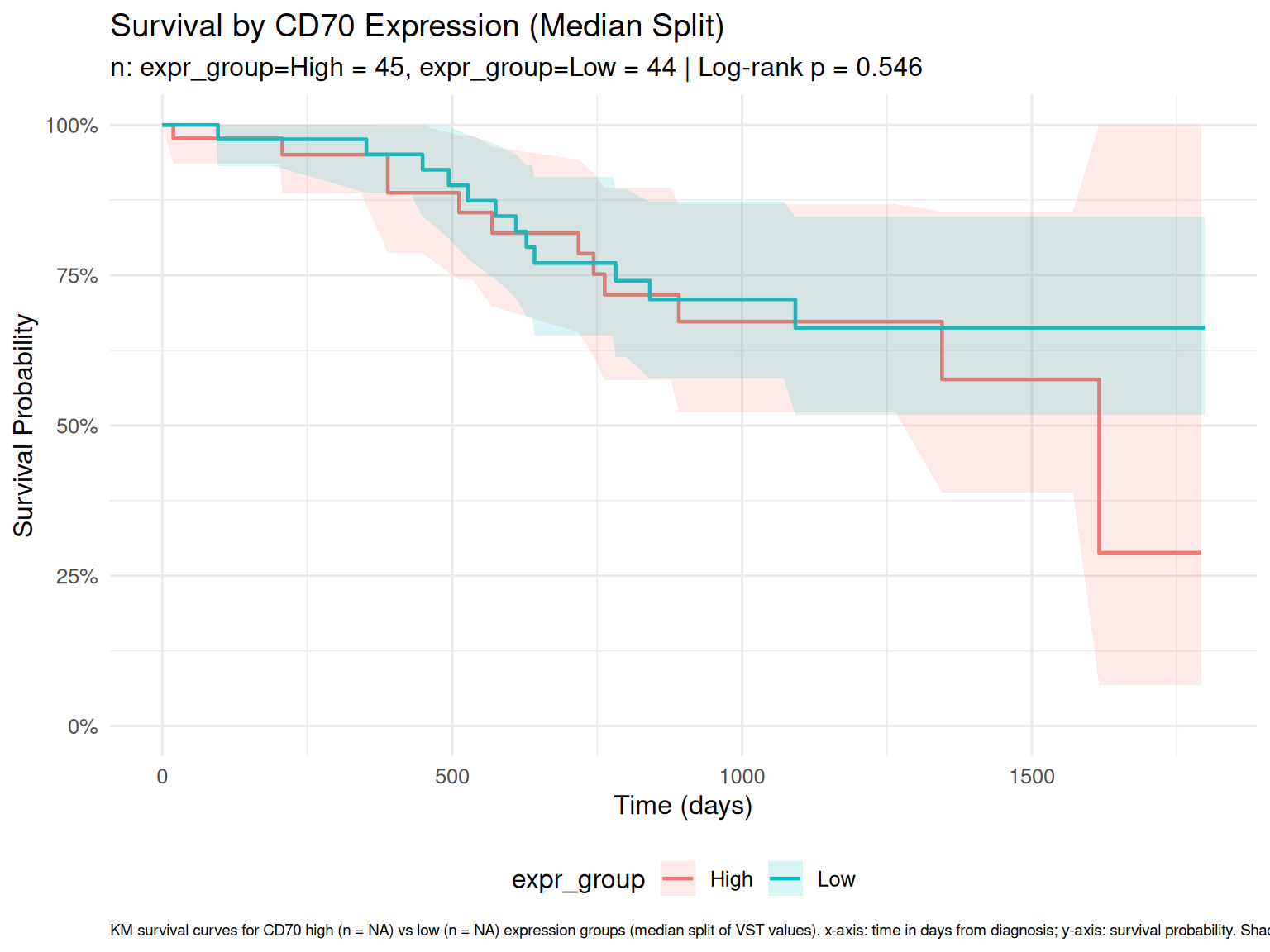
Survival by Expression
Patients split at median CD70 expression into High/Low groups.
Expression by Cytogenetic Subtype
Boxplots comparing expression levels across FISH-defined cytogenetic marker groups (t(4;14), t(14;16), del(17p)).
Data not available: Target
vig_gene_expr_subtype returned NULL. See
Data Sources.
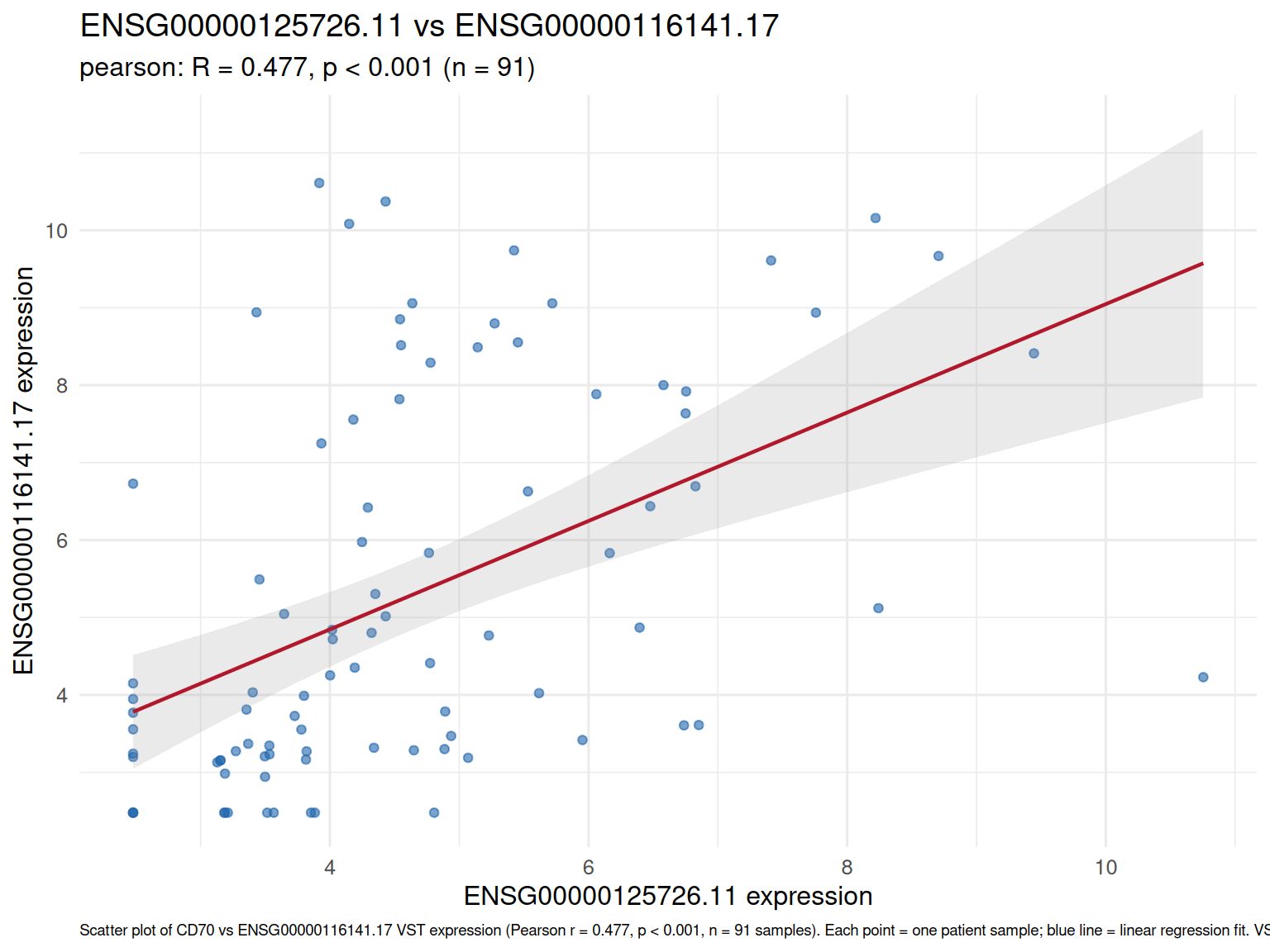
Gene-Gene Correlations
Top correlated genes with CD70 (Pearson, VST expression).
Top 10 genes most correlated with CD70 (Pearson r, VST-normalized expression, n = 91 samples). Strongest correlation: ENSG00000116141.17 (r = 0.477). estimate = Pearson correlation coefficient; p_value = unadjusted; padj = BH-corrected. Candidates: top 500 most variable genes. See scatter plot below for visualization of the top correlation.
|
gene
|
estimate
|
p_value
|
padj
|
|
ENSG00000116141.17
|
0.477
|
1.80e-06
|
0.000411
|
|
ENSG00000157985.19
|
0.471
|
2.40e-06
|
0.000411
|
|
ENSG00000110092.4
|
-0.470
|
2.70e-06
|
0.000411
|
|
ENSG00000185630.19
|
0.466
|
3.30e-06
|
0.000411
|
|
ENSG00000154277.13
|
0.456
|
5.60e-06
|
0.000557
|
|
ENSG00000183580.10
|
0.441
|
1.22e-05
|
0.001015
|
|
ENSG00000204219.11
|
0.429
|
2.21e-05
|
0.001580
|
|
ENSG00000118971.9
|
0.425
|
2.71e-05
|
0.001696
|
|
ENSG00000148468.17
|
0.422
|
3.12e-05
|
0.001731
|
|
ENSG00000211677.2
|
0.403
|
7.56e-05
|
0.003780
|
Data Sources
Results in this vignette are derived from the MMRF CoMMpass study (MMRF-COMMPASS, ~1,143 patients), downloaded via TCGAbiolinks. The pipeline runs with a configurable sample_limit (default 200; CI uses 20).
For full citations, data access tiers, and the distinction between pipeline data and synthetic test data, see the Data Sources vignette.
Recent Changes
Recent project commits with lines added, files changed, and change categories.
Last 20 project commits with change statistics. Date = commit date; Type = conventional-commit prefix (feat/fix/docs/ci/refactor/test/chore). Files = number of files modified; +Lines/-Lines = lines added/removed. Source: git log –numstat. See changes-by-type table for aggregate breakdown.
|
date
|
type
|
summary
|
n_files
|
lines_added
|
lines_removed
|
file_categories
|
|
2026-03-14
|
Bug Fix
|
fix(pipeline): Fix 11 NULL targets — DE condition, ID matching, consensus type
|
41
|
146
|
47
|
Other, R Source
|
|
2026-03-14
|
Bug Fix
|
fix(cachix): Remove –watch-mode auto flag (already default)
|
1
|
1
|
1
|
Other
|
|
2026-03-14
|
Bug Fix
|
fix(pipeline): Fix 3 NULL-target bugs, auto-generate package.nix (#93)
|
87
|
235
|
80
|
Config, Docs, Other, R Source
|
|
2026-03-14
|
Bug Fix
|
fix(nix): Fix cachix signing key, rebuild Bioconductor-dependent targets
|
2
|
0
|
0
|
Other
|
|
2026-03-14
|
New Feature
|
feat(captions): Add dynamic captions to 34 table/plot targets
|
22
|
579
|
89
|
Other, R Source
|
|
2026-03-14
|
Bug Fix
|
fix(vignettes): Enforce zero-computation rule — 22 violations → 0
|
32
|
360
|
764
|
Other, R Source, Vignettes
|
|
2026-03-13
|
Bug Fix
|
fix(vignettes): Convert kable RDS to data.frames, fix telemetry eval guards
|
18
|
8
|
2
|
Other, Vignettes
|
|
2026-03-13
|
Bug Fix
|
fix(ci): Save data frames (not DT widgets) to RDS for Nix portability
|
3
|
0
|
0
|
Other
|
|
2026-03-13
|
Bug Fix
|
fix(vignettes): Use Quarto #| eval syntax for pkgdown-banner chunks
|
11
|
44
|
11
|
Vignettes
|
|
2026-03-13
|
Refactoring
|
refactor(targets): Move Bioconductor packages to per-target declarations
|
11
|
35
|
17
|
Other, R Source, Vignettes
|
|
2026-03-13
|
New Feature
|
feat(vignettes): Add code provenance, kable→DT conversion, caption compliance
|
35
|
1004
|
437
|
CI/CD, Other, R Source, Vignettes
|
|
2026-03-13
|
Bug Fix
|
fix(vignettes): Skip NULL RDS in safe_tar_read, return invisible(NULL)
|
11
|
22
|
22
|
Vignettes
|
|
2026-03-13
|
Bug Fix
|
fix(glossary): Prevent double DT::datatable() wrapping in glossary-table chunk
|
1
|
3
|
1
|
Vignettes
|
|
2026-03-13
|
CI/CD
|
ci: Show quarto errors with quiet=FALSE, render individual vignettes in diagnostic
|
1
|
20
|
6
|
CI/CD
|
|
2026-03-13
|
CI/CD
|
ci: Add verbose quarto error diagnostics on build failure
|
1
|
14
|
1
|
CI/CD
|
|
2026-03-13
|
Bug Fix
|
fix(vignettes): Strip Nix paths from DT widgets, auto-wrap data frames
|
25
|
66
|
28
|
CI/CD, Other, Vignettes
|
|
2026-03-13
|
CI/CD
|
ci: Add diagnostic quarto render step to debug build failure
|
1
|
17
|
0
|
CI/CD
|
|
2026-03-13
|
Bug Fix
|
fix(vignettes): Revert safe_tar_read placeholder, guard gene-report
|
11
|
12
|
56
|
Vignettes
|
|
2026-03-13
|
Maintenance
|
chore: Export vig_count_distribution_plot as ggplot RDS (513KB)
|
1
|
0
|
0
|
Other
|
|
2026-03-13
|
Bug Fix
|
fix(vignettes): Enable code eval in CI with RDS fallback
|
80
|
113
|
74
|
CI/CD, Other, R Source, Vignettes
|
Reproducibility
Session Info (click to expand)
Show code
sessionInfo()
#> R version 4.5.3 (2026-03-11)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.3 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> loaded via a namespace (and not attached):
#> [1] Matrix_1.7-4 base64url_1.4 gtable_0.3.6
#> [4] jsonlite_2.0.0 dplyr_1.2.0 compiler_4.5.3
#> [7] tidyselect_1.2.1 callr_3.7.6 splines_4.5.3
#> [10] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0
#> [13] lattice_0.22-9 ggplot2_4.0.2 R6_2.6.1
#> [16] labeling_0.4.3 generics_0.1.4 igraph_2.2.2
#> [19] knitr_1.51 backports_1.5.0 targets_1.12.0
#> [22] tibble_3.3.1 pillar_1.11.1 RColorBrewer_1.1-3
#> [25] rlang_1.1.7 xfun_0.57 S7_0.2.1
#> [28] otel_0.2.0 cli_3.6.5 mgcv_1.9-4
#> [31] withr_3.0.2 magrittr_2.0.4 ps_1.9.1
#> [34] digest_0.6.39 grid_4.5.3 processx_3.8.6
#> [37] nlme_3.1-168 secretbase_1.2.0 lifecycle_1.0.5
#> [40] prettyunits_1.2.0 vctrs_0.7.2 evaluate_1.0.5
#> [43] glue_1.8.0 data.table_1.18.2.1 farver_2.1.2
#> [46] codetools_0.2-20 rmarkdown_2.30 tools_4.5.3
#> [49] pkgconfig_2.0.3 htmltools_0.5.9

