flowchart LR
subgraph GDC["GDC Open Access"]
R["RNA-seq<br/>STAR Counts"]
C["Clinical<br/>Metadata"]
T["Treatment<br/>Records"]
end
subgraph Clean["Data Cleaning"]
SE["Summarized<br/>Experiment"]
CD["Clinical<br/>DataFrame"]
TD["Treatment<br/>DataFrame"]
end
subgraph Analysis
DE["Differential<br/>Expression"]
KM["Survival<br/>Analysis"]
PA["Pathway<br/>Enrichment"]
DAG["Causal<br/>DAGs"]
end
R --> SE
C --> CD
T --> TD
SE --> DE
CD --> KM
TD --> KM
DE --> PA
SE --> KM
CD --> DAG
style GDC fill:#e8f5e9,stroke:#4CAF50
style Clean fill:#e3f2fd,stroke:#2196F3
style Analysis fill:#fce4ec,stroke:#F44336
Online documentation
This vignette displays pre-computed results. Run the targets pipeline locally for interactive analysis.
Overview
See the Glossary for term definitions (read count, library size, gene detection, outlier, etc.).
- RNA-seq and clinical data from GDC via the
targetspipeline - Open Access RNA-seq from the GDC S3 bucket (
s3://gdc-mmrf-commpass-phs000748-2-open/) - Clinical metadata via
TCGAbiolinks::GDCquery_clinic()
Data Flow
Pipeline Configuration
Current pipeline settings including sample limit, random seed, and data paths.
The pipeline downloads RNA-seq and clinical data from the GDC portal (https://portal.gdc.cancer.gov/projects/MMRF-COMMPASS) for the MMRF-COMMPASS project.
Sample limit: 200 patients (GDC has ~900; subsetting speeds local development). In CI, this is capped at 20 samples (see R/01_data_acquisition.R) to keep build times under 10 minutes.
Random seed: 42 — ensures reproducible patient selection when random_sample = TRUE.
Data directory: data | Results directory: results
Note: This vignette was built with
sample_limit = 200patients. Numbers below reflect this subset, not the full ~900-patient CoMMpass cohort. For pipeline execution details, see issue #46 (telemetry vignette).
RNA-seq Data
RNA-seq gene expression data is downloaded from GDC as a SummarizedExperiment object containing STAR-Counts.
RNA-seq Data Summary
- Samples: 100
- Genes: 60,660
- Total counts: 6,045,546,169 (sum of all read counts across all genes and samples)
- Median counts per sample: 58,030,298 (median library size)
- Sparsity (% zero entries): 49.1%
| Sample | Total_Counts | Genes_Detected | Median_Count | Max_Count | |
|---|---|---|---|---|---|
| MMRF_1618_1_BM_CD138pos | MMRF_1618_1_BM_CD138pos | 37391891 | 27609 | 0 | 4187802 |
| MMRF_2229_1_BM_CD138pos | MMRF_2229_1_BM_CD138pos | 51745321 | 28905 | 0 | 9973468 |
| MMRF_2143_1_BM_CD138pos | MMRF_2143_1_BM_CD138pos | 59392637 | 28249 | 0 | 7755820 |
| MMRF_2054_1_BM_CD138pos | MMRF_2054_1_BM_CD138pos | 63680199 | 32356 | 1 | 6909884 |
| MMRF_1700_2_BM_CD138pos | MMRF_1700_2_BM_CD138pos | 96656035 | 37358 | 3 | 7104305 |
| MMRF_1871_1_BM_CD138pos | MMRF_1871_1_BM_CD138pos | 46761239 | 28751 | 0 | 4537227 |
| MMRF_1293_1_BM_CD138pos | MMRF_1293_1_BM_CD138pos | 50232734 | 30109 | 0 | 2463705 |
| MMRF_1715_1_BM_CD138pos | MMRF_1715_1_BM_CD138pos | 58060644 | 30504 | 1 | 4930116 |
| MMRF_1637_1_BM_CD138pos | MMRF_1637_1_BM_CD138pos | 41424544 | 30874 | 1 | 4524926 |
| MMRF_1543_1_BM_CD138pos | MMRF_1543_1_BM_CD138pos | 44701993 | 27289 | 0 | 4206708 |
| MMRF_1698_1_BM_CD138pos | MMRF_1698_1_BM_CD138pos | 46684067 | 25832 | 0 | 14630370 |
| MMRF_1716_1_BM_CD138pos | MMRF_1716_1_BM_CD138pos | 46903826 | 27259 | 0 | 7237680 |
| MMRF_2341_1_BM_CD138pos | MMRF_2341_1_BM_CD138pos | 71582628 | 33440 | 1 | 3881877 |
| MMRF_2085_3_BM_CD138pos | MMRF_2085_3_BM_CD138pos | 101561094 | 34138 | 1 | 5931977 |
| MMRF_2422_1_BM_CD138pos | MMRF_2422_1_BM_CD138pos | 49591878 | 28663 | 0 | 12074308 |
| MMRF_2429_1_BM_CD138pos | MMRF_2429_1_BM_CD138pos | 44521218 | 29397 | 0 | 6632843 |
| MMRF_2245_1_BM_CD138pos | MMRF_2245_1_BM_CD138pos | 55815128 | 30949 | 1 | 8274019 |
| MMRF_2043_1_BM_CD138pos | MMRF_2043_1_BM_CD138pos | 70996601 | 30698 | 1 | 9214029 |
| MMRF_2401_2_BM_CD138pos | MMRF_2401_2_BM_CD138pos | 76584241 | 32974 | 1 | 9970865 |
| MMRF_2098_1_BM_CD138pos | MMRF_2098_1_BM_CD138pos | 57538387 | 29719 | 0 | 13565805 |
| MMRF_2238_1_BM_CD138pos | MMRF_2238_1_BM_CD138pos | 66314228 | 33476 | 1 | 9190976 |
| MMRF_1436_1_BM_CD138pos | MMRF_1436_1_BM_CD138pos | 57999952 | 29968 | 0 | 12652198 |
| MMRF_1108_1_BM_CD138pos | MMRF_1108_1_BM_CD138pos | 42289892 | 28079 | 0 | 6358971 |
| MMRF_1856_1_BM_CD138pos | MMRF_1856_1_BM_CD138pos | 48755441 | 28313 | 0 | 7487960 |
| MMRF_2746_1_BM_CD138pos | MMRF_2746_1_BM_CD138pos | 74850398 | 35919 | 2 | 10146793 |
| MMRF_1137_4_BM_CD138pos | MMRF_1137_4_BM_CD138pos | 55587700 | 32731 | 1 | 5680921 |
| MMRF_1361_1_BM_CD138pos | MMRF_1361_1_BM_CD138pos | 76274233 | 34427 | 1 | 7338549 |
| MMRF_1048_1_BM_CD138pos | MMRF_1048_1_BM_CD138pos | 46040142 | 28127 | 0 | 4406314 |
| MMRF_2828_1_BM_CD138pos | MMRF_2828_1_BM_CD138pos | 68349206 | 27973 | 0 | 16875619 |
| MMRF_2601_1_BM_CD138pos | MMRF_2601_1_BM_CD138pos | 39797530 | 28415 | 0 | 5453712 |
| MMRF_2232_1_BM_CD138pos | MMRF_2232_1_BM_CD138pos | 81630421 | 31401 | 1 | 17517416 |
| MMRF_2253_1_BM_CD138pos | MMRF_2253_1_BM_CD138pos | 63958386 | 32111 | 1 | 13127382 |
| MMRF_1824_1_BM_CD138pos | MMRF_1824_1_BM_CD138pos | 37401747 | 28419 | 0 | 6215715 |
| MMRF_2197_1_BM_CD138pos | MMRF_2197_1_BM_CD138pos | 65930119 | 33595 | 1 | 8263777 |
| MMRF_2664_1_BM_CD138pos | MMRF_2664_1_BM_CD138pos | 64857697 | 31545 | 1 | 3502164 |
| MMRF_1819_1_BM_CD138pos | MMRF_1819_1_BM_CD138pos | 49401746 | 28052 | 0 | 8330859 |
| MMRF_2716_1_BM_CD138pos | MMRF_2716_1_BM_CD138pos | 91426018 | 34504 | 1 | 19264905 |
| MMRF_1617_1_BM_CD138pos | MMRF_1617_1_BM_CD138pos | 37826511 | 27819 | 0 | 3335089 |
| MMRF_2199_1_BM_CD138pos | MMRF_2199_1_BM_CD138pos | 52651580 | 28971 | 0 | 8173315 |
| MMRF_1223_2_BM_CD138pos | MMRF_1223_2_BM_CD138pos | 64253003 | 30235 | 0 | 12418148 |
| MMRF_1216_1_BM_CD138pos | MMRF_1216_1_BM_CD138pos | 54430074 | 31804 | 1 | 9765806 |
| MMRF_2705_1_BM_CD138pos | MMRF_2705_1_BM_CD138pos | 149432793 | 35625 | 2 | 15342982 |
| MMRF_2257_1_BM_CD138pos | MMRF_2257_1_BM_CD138pos | 52847814 | 32503 | 1 | 5837429 |
| MMRF_2815_1_BM_CD138pos | MMRF_2815_1_BM_CD138pos | 54076776 | 31299 | 1 | 14840619 |
| MMRF_2636_1_BM_CD138pos | MMRF_2636_1_BM_CD138pos | 79930092 | 31067 | 1 | 15922682 |
| MMRF_1651_1_BM_CD138pos | MMRF_1651_1_BM_CD138pos | 38531104 | 29803 | 0 | 4537182 |
| MMRF_1991_1_BM_CD138pos | MMRF_1991_1_BM_CD138pos | 69001805 | 31154 | 1 | 10006934 |
| MMRF_1502_1_BM_CD138pos | MMRF_1502_1_BM_CD138pos | 60836136 | 31923 | 1 | 7502143 |
| MMRF_1049_4_BM_CD138pos | MMRF_1049_4_BM_CD138pos | 67752904 | 31171 | 1 | 6880242 |
| MMRF_1250_1_BM_CD138pos | MMRF_1250_1_BM_CD138pos | 41495680 | 30909 | 1 | 3727969 |
| MMRF_1496_1_PB_CD138pos | MMRF_1496_1_PB_CD138pos | 79637573 | 32560 | 1 | 7554268 |
| MMRF_1496_1_BM_CD138pos | MMRF_1496_1_BM_CD138pos | 149615704 | 34145 | 1 | 8690744 |
| MMRF_1235_1_BM_CD138pos | MMRF_1235_1_BM_CD138pos | 60749662 | 31612 | 1 | 10475083 |
| MMRF_1978_1_BM_CD138pos | MMRF_1978_1_BM_CD138pos | 58788425 | 31621 | 1 | 8629307 |
| MMRF_2089_3_BM_CD138pos | MMRF_2089_3_BM_CD138pos | 87760456 | 34575 | 1 | 6546283 |
| MMRF_1900_1_BM_CD138pos | MMRF_1900_1_BM_CD138pos | 52217137 | 32128 | 1 | 6369728 |
| MMRF_2224_1_BM_CD138pos | MMRF_2224_1_BM_CD138pos | 54403435 | 31139 | 1 | 2535879 |
| MMRF_2458_1_BM_CD138pos | MMRF_2458_1_BM_CD138pos | 56294068 | 29650 | 0 | 7076758 |
| MMRF_1092_1_BM_CD138pos | MMRF_1092_1_BM_CD138pos | 36748951 | 29135 | 0 | 4034219 |
| MMRF_2822_1_BM_CD138pos | MMRF_2822_1_BM_CD138pos | 60054775 | 32643 | 1 | 10858052 |
| MMRF_1538_1_BM_CD138pos | MMRF_1538_1_BM_CD138pos | 61613394 | 29038 | 0 | 7612687 |
| MMRF_1992_1_PB_CD138pos | MMRF_1992_1_PB_CD138pos | 40521003 | 26760 | 0 | 6365931 |
| MMRF_2729_1_BM_CD138pos | MMRF_2729_1_BM_CD138pos | 94208666 | 35598 | 2 | 12749799 |
| MMRF_1625_1_BM_CD138pos | MMRF_1625_1_BM_CD138pos | 38223103 | 25966 | 0 | 6474610 |
| MMRF_2107_1_BM_CD138pos | MMRF_2107_1_BM_CD138pos | 63872226 | 31475 | 1 | 12848329 |
| MMRF_2055_1_BM_CD138pos | MMRF_2055_1_BM_CD138pos | 41457997 | 29706 | 0 | 5020097 |
| MMRF_1944_1_BM_CD138pos | MMRF_1944_1_BM_CD138pos | 47103245 | 30647 | 1 | 5660731 |
| MMRF_1110_2_BM_CD138pos | MMRF_1110_2_BM_CD138pos | 77343762 | 30631 | 1 | 6714330 |
| MMRF_2228_1_BM_CD138pos | MMRF_2228_1_BM_CD138pos | 68086550 | 33922 | 1 | 7433572 |
| MMRF_1129_1_BM_CD138pos | MMRF_1129_1_BM_CD138pos | 63899643 | 31634 | 1 | 12076907 |
| MMRF_1783_2_BM_CD138pos | MMRF_1783_2_BM_CD138pos | 82754792 | 35563 | 2 | 16490062 |
| MMRF_1783_1_BM_CD138pos | MMRF_1783_1_BM_CD138pos | 50776425 | 30821 | 1 | 15358485 |
| MMRF_2832_1_BM_CD138pos | MMRF_2832_1_BM_CD138pos | 61360677 | 28685 | 0 | 5405622 |
| MMRF_1778_1_PB_CD138pos | MMRF_1778_1_PB_CD138pos | 67550482 | 34614 | 2 | 9375160 |
| MMRF_1267_1_BM_CD138pos | MMRF_1267_1_BM_CD138pos | 30377770 | 27628 | 0 | 7257740 |
| MMRF_2843_1_BM_CD138pos | MMRF_2843_1_BM_CD138pos | 77359789 | 31052 | 1 | 2570969 |
| MMRF_2047_1_BM_CD138pos | MMRF_2047_1_BM_CD138pos | 39751015 | 29294 | 0 | 7309250 |
| MMRF_1613_1_BM_CD138pos | MMRF_1613_1_BM_CD138pos | 44241364 | 29485 | 0 | 6563231 |
| MMRF_1445_1_BM_CD138pos | MMRF_1445_1_BM_CD138pos | 53144209 | 30277 | 0 | 9002435 |
| MMRF_1556_1_BM_CD138pos | MMRF_1556_1_BM_CD138pos | 74700167 | 32015 | 1 | 9120954 |
| MMRF_2827_1_BM_CD138pos | MMRF_2827_1_BM_CD138pos | 61996840 | 32601 | 1 | 6359115 |
| MMRF_1252_1_BM_CD138pos | MMRF_1252_1_BM_CD138pos | 22018481 | 26953 | 0 | 4914261 |
| MMRF_1533_2_BM_CD138pos | MMRF_1533_2_BM_CD138pos | 46280409 | 30627 | 1 | 4543360 |
| MMRF_1912_1_PB_CD138pos | MMRF_1912_1_PB_CD138pos | 55599192 | 32521 | 1 | 6738655 |
| MMRF_2153_1_BM_CD138pos | MMRF_2153_1_BM_CD138pos | 62246722 | 32147 | 1 | 7229388 |
| MMRF_1356_1_BM_CD138pos | MMRF_1356_1_BM_CD138pos | 48785009 | 30935 | 1 | 8235691 |
| MMRF_1033_1_BM_CD138pos | MMRF_1033_1_BM_CD138pos | 52817165 | 28544 | 0 | 16876687 |
| MMRF_1079_1_BM_CD138pos | MMRF_1079_1_BM_CD138pos | 81118832 | 32913 | 1 | 6687030 |
| MMRF_1401_2_BM_CD138pos | MMRF_1401_2_BM_CD138pos | 67018253 | 27599 | 0 | 8923547 |
| MMRF_1621_1_BM_CD138pos | MMRF_1621_1_BM_CD138pos | 46791382 | 29873 | 0 | 6646887 |
| MMRF_2225_1_BM_CD138pos | MMRF_2225_1_BM_CD138pos | 58724738 | 32595 | 1 | 2236229 |
| MMRF_1501_1_BM_CD138pos | MMRF_1501_1_BM_CD138pos | 60234555 | 32331 | 1 | 5609859 |
| MMRF_2763_1_BM_CD138pos | MMRF_2763_1_BM_CD138pos | 48752427 | 29503 | 0 | 5358227 |
| MMRF_2150_1_BM_CD138pos | MMRF_2150_1_BM_CD138pos | 55307603 | 27982 | 0 | 9654610 |
| MMRF_1730_1_BM_CD138pos | MMRF_1730_1_BM_CD138pos | 60872279 | 29038 | 0 | 10674765 |
| MMRF_2675_1_BM_CD138pos | MMRF_2675_1_BM_CD138pos | 73461172 | 32431 | 1 | 8861444 |
| MMRF_1690_1_BM_CD138pos | MMRF_1690_1_BM_CD138pos | 56634991 | 29767 | 0 | 8512297 |
| MMRF_1932_1_BM_CD138pos | MMRF_1932_1_BM_CD138pos | 58691302 | 32437 | 1 | 2462320 |
| MMRF_2072_1_BM_CD138pos | MMRF_2072_1_BM_CD138pos | 78601932 | 34471 | 2 | 5425361 |
| MMRF_1462_3_BM_CD138pos | MMRF_1462_3_BM_CD138pos | 49218987 | 29169 | 0 | 6212039 |
Generating code
{
if (!file.exists(raw_rnaseq))
return(NULL)
se <- readRDS(raw_rnaseq)
if (!inherits(se, "SummarizedExperiment"))
return(NULL)
counts <- get_counts_assay(se)
log_counts <- log10(counts + 1)
mean_expr <- rowMeans(log_counts)
gene_meta <- as.data.frame(SummarizedExperiment::rowData(se))
has_biotype <- "gene_type" %in% names(gene_meta)
if (has_biotype) {
gene_meta$biotype_group <- dplyr::case_when(gene_meta$gene_type ==
"protein_coding" ~ "protein-coding", gene_meta$gene_type %in%
c("lncRNA", "processed_pseudogene", "unprocessed_pseudogene",
"transcribed_unprocessed_pseudogene", "transcribed_processed_pseudogene") ~
"lncRNA / pseudogene", TRUE ~ "other (miRNA, snoRNA, etc.)")
plot_df <- data.frame(mean_expr = mean_expr, biotype = gene_meta$biotype_group,
stringsAsFactors = FALSE)
biotype_colors <- c(`protein-coding` = "#0066CC", `lncRNA / pseudogene` = "#DC3545",
`other (miRNA, snoRNA, etc.)` = "#6C757D")
p <- ggplot2::ggplot(plot_df, ggplot2::aes(x = mean_expr,
fill = biotype)) + ggplot2::geom_histogram(bins = 50,
alpha = 0.6, position = "identity") + ggplot2::scale_fill_manual(values = biotype_colors) +
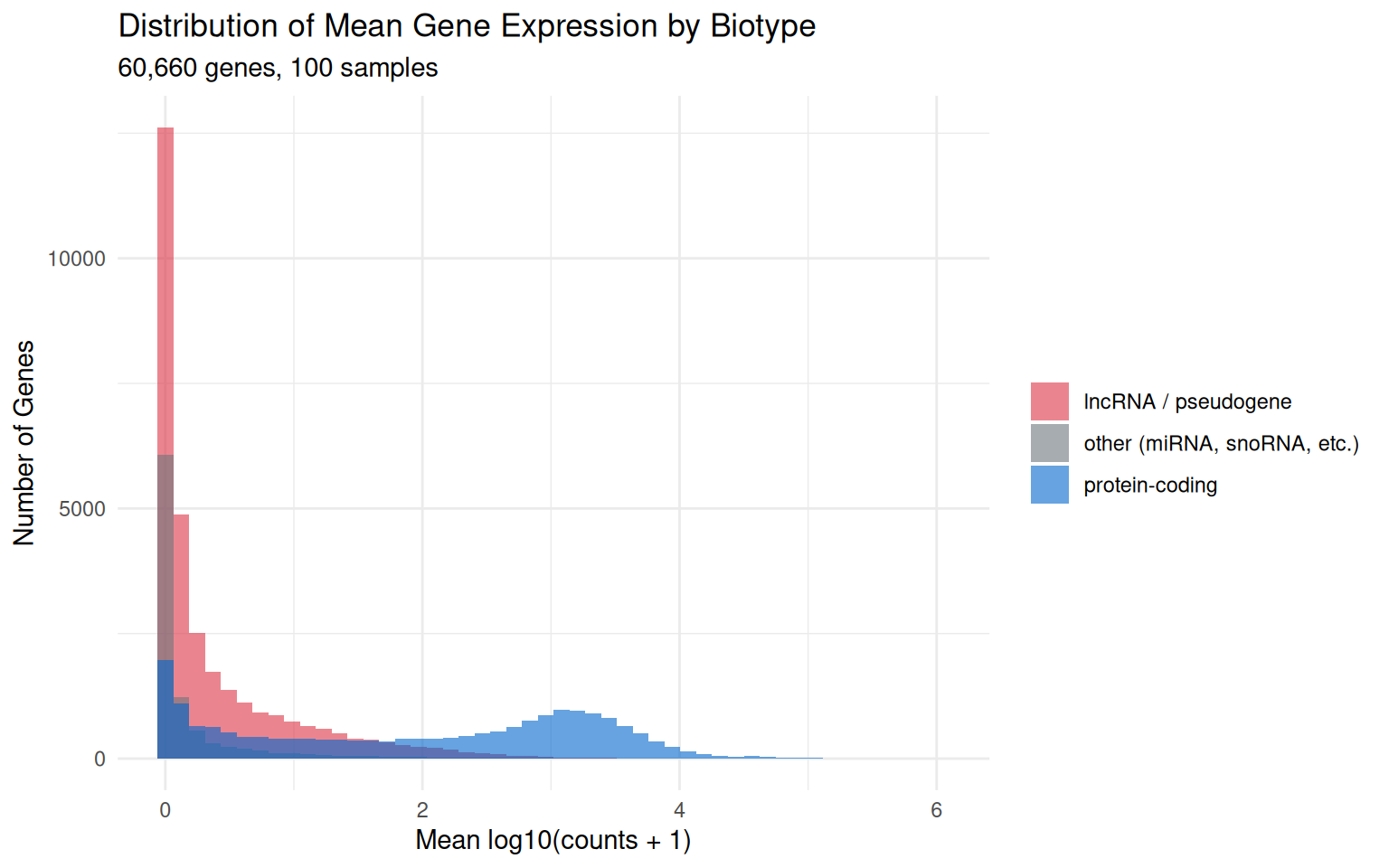
ggplot2::labs(title = "Distribution of Mean Gene Expression by Biotype",
subtitle = paste0(format(nrow(counts), big.mark = ","),
" genes, ", ncol(counts), " samples"), x = "Mean log10(counts + 1)",
y = "Number of Genes", fill = NULL) + ggplot2::theme_minimal()
}
else {
plot_df <- data.frame(mean_expr = mean_expr)
p <- ggplot2::ggplot(plot_df, ggplot2::aes(x = mean_expr)) +
ggplot2::geom_histogram(bins = 50, fill = "steelblue",
alpha = 0.7) + ggplot2::labs(title = "Distribution of Mean Gene Expression",
subtitle = paste0(format(nrow(counts), big.mark = ","),
" genes, ", ncol(counts), " samples"), x = "Mean log10(counts + 1)",
y = "Number of Genes") + ggplot2::theme_minimal()
}
p
}
Clinical Data
Clinical metadata from GDC provides patient demographics, disease characteristics, and outcomes. See the data dictionary for variable definitions and the glossary for units.
| Column | Type | Non_NA | Pct_Complete | N_Unique | Example | |
|---|---|---|---|---|---|---|
| 1 | project | character | 995 | 100% | 1 | MMRF-COMMPASS |
| 2 | submitter_id | character | 995 | 100% | 995 | MMRF_1016, MMRF_1020, MMRF_1021 |
| 3 | submitter_id.1 | character | 995 | 100% | 995 | MMRF_1016, MMRF_1020, MMRF_1021 |
| 4 | submitter_sample_ids | character | 995 | 100% | 995 | MMRF_1016_1_BM_CD138pos,MMRF_1016_1_PB_WBC, MMRF_1020_3_BM_CD138pos,MMRF_1020_3_PB_Whole, MMRF_1021_1_BM_CD138pos,MMRF_1021_2_PB_CD3pos |
| 5 | submitter_id.2 | character | 995 | 100% | 995 | MMRF_10161, MMRF_10201, MMRF_10211 |
| 6 | age_at_diagnosis_days | character | 995 | 100% | 927 | NA, 17034, 18560 |
| 7 | race | character | 995 | 100% | 6 | white, not reported, black or african american |
| 8 | gender | character | 995 | 100% | 2 | male, female |
| 9 | ethnicity | character | 995 | 100% | 3 | not hispanic or latino, not reported, hispanic or latino |
| 10 | age_at_index_days | integer | 995 | 100% | 59 | min=27 med=63 max=89 |
| 11 | days_to_last_known_disease_status_days | character | 995 | 100% | 693 | 1, 997, 617 |
| 12 | days_to_best_overall_response | character | 995 | 100% | 1 | NA |
| 13 | days_to_diagnosis | character | 995 | 100% | 1 | NA |
| 14 | days_to_last_follow_up_days | character | 995 | 100% | 693 | 1, 997, 617 |
| 15 | year_of_diagnosis | character | 995 | 100% | 1 | NA |
| 16 | days_to_recurrence | character | 995 | 100% | 1 | NA |
| 17 | days_to_birth | integer | 953 | 96% | 926 | min=-32800 med=-23200 max=-10100 |
| 18 | year_of_birth | logical | 0 | 0% | 0 | (all NA) |
| 19 | days_to_death_days | integer | 191 | 19% | 176 | min=8 med=475 max=1750 |
| 20 | year_of_death | logical | 0 | 0% | 0 | (all NA) |
| 21 | last_known_disease_status | character | 995 | 100% | 1 | Unknown tumor status |
| 22 | cause_of_death | character | 188 | 19% | 2 | Cancer Related, Not Cancer Related |
| 23 | vital_status | character | 995 | 100% | 2 | Alive, Dead |
| 24 | primary_site | character | 995 | 100% | 1 | Hematopoietic and reticuloendothelial systems |
| 25 | irs_stage | character | 995 | 100% | 1 | NA |
| 26 | iss_stage | character | 995 | 100% | 4 | II, I, III |
| 27 | ajcc_pathologic_stage | character | 995 | 100% | 1 | NA |
| 28 | ann_arbor_clinical_stage | character | 995 | 100% | 1 | NA |
| 29 | enneking_msts_stage | character | 995 | 100% | 1 | NA |
| 30 | inrg_stage | character | 995 | 100% | 1 | NA |
| 31 | tissue_or_organ_of_origin | character | 995 | 100% | 1 | Bone marrow |
| 32 | cog_liver_stage | character | 995 | 100% | 1 | NA |
| 33 | inpc_grade | character | 995 | 100% | 1 | NA |
| 34 | wilms_tumor_histologic_subtype | character | 995 | 100% | 1 | NA |
| 35 | classification_of_tumor | character | 995 | 100% | 1 | NA |
| 36 | cog_renal_stage | character | 995 | 100% | 1 | NA |
| 37 | figo_stage | character | 995 | 100% | 1 | NA |
| 38 | inss_stage | character | 995 | 100% | 1 | NA |
| 39 | tumor_confined_to_organ_of_origin | character | 995 | 100% | 1 | NA |
| 40 | primary_diagnosis | character | 995 | 100% | 1 | Multiple myeloma |
| 41 | ajcc_clinical_stage | character | 995 | 100% | 1 | NA |
| 42 | metastasis_at_diagnosis | character | 995 | 100% | 1 | NA |
| 43 | enneking_msts_tumor_site | character | 995 | 100% | 1 | NA |
| 44 | ann_arbor_pathologic_stage | character | 995 | 100% | 1 | NA |
| 45 | method_of_diagnosis | character | 995 | 100% | 1 | NA |
| 46 | diagnosis_id | character | 995 | 100% | 995 | 0073d350-15ca-4e89-9326-d95543cf778c, 00a8b803-c44b-41ba-9d44-79c1b87a74aa, 00fe922e-5ecb-4936-a91d-6680d3f393c6 |
| 47 | site_of_resection_or_biopsy | character | 995 | 100% | 1 | Bone marrow |
| 48 | first_symptom_prior_to_diagnosis | character | 995 | 100% | 1 | NA |
| 49 | tumor_grade | character | 995 | 100% | 1 | Unknown |
| 50 | enneking_msts_grade | character | 995 | 100% | 1 | NA |
| 51 | disease_type | character | 995 | 100% | 1 | Plasma Cell Tumors |
| 52 | created_datetime | character | 995 | 100% | 1 | 2018-07-10T14:08:13.021252-05:00 |
| 53 | enneking_msts_metastasis | character | 995 | 100% | 1 | NA |
| 54 | esophageal_columnar_dysplasia_degree | character | 995 | 100% | 1 | NA |
| 55 | child_pugh_classification | character | 995 | 100% | 1 | NA |
| 56 | state | character | 995 | 100% | 1 | released |
| 57 | prior_treatment | character | 995 | 100% | 1 | NA |
| 58 | cog_rhabdomyosarcoma_risk_group | character | 995 | 100% | 1 | NA |
| 59 | ajcc_pathologic_t | character | 995 | 100% | 1 | NA |
| 60 | morphology | character | 995 | 100% | 1 | 9732/3 |
| 61 | ajcc_pathologic_n | character | 995 | 100% | 1 | NA |
| 62 | ajcc_pathologic_m | character | 995 | 100% | 1 | NA |
| 63 | irs_group | character | 995 | 100% | 1 | NA |
| 64 | medulloblastoma_molecular_classification | character | 995 | 100% | 1 | NA |
| 65 | residual_disease | character | 995 | 100% | 1 | NA |
| 66 | ann_arbor_b_symptoms | character | 995 | 100% | 1 | NA |
| 67 | icd_10_code | character | 995 | 100% | 1 | NA |
| 68 | synchronous_malignancy | character | 995 | 100% | 1 | NA |
| 69 | burkitt_lymphoma_clinical_variant | character | 995 | 100% | 1 | NA |
| 70 | supratentorial_localization | character | 995 | 100% | 1 | NA |
| 71 | ishak_fibrosis_score | character | 995 | 100% | 1 | NA |
| 72 | goblet_cells_columnar_mucosa_present | character | 995 | 100% | 1 | NA |
| 73 | laterality | character | 995 | 100% | 1 | NA |
| 74 | cog_neuroblastoma_risk_group | character | 995 | 100% | 1 | NA |
| 75 | updated_datetime | character | 995 | 100% | 2 | 2019-06-24T08:07:19.797044-05:00, 2019-08-21T12:47:37.999949-05:00 |
| 76 | prior_malignancy | character | 995 | 100% | 1 | NA |
| 77 | best_overall_response | character | 995 | 100% | 1 | NA |
| 78 | ann_arbor_extranodal_involvement | character | 995 | 100% | 1 | NA |
| 79 | mitosis_karyorrhexis_index | character | 995 | 100% | 1 | NA |
| 80 | ajcc_staging_system_edition | character | 995 | 100% | 1 | NA |
| 81 | esophageal_columnar_metaplasia_present | character | 995 | 100% | 1 | NA |
| 82 | ajcc_clinical_m | character | 995 | 100% | 1 | NA |
| 83 | ajcc_clinical_n | character | 995 | 100% | 1 | NA |
| 84 | ajcc_clinical_t | character | 995 | 100% | 1 | NA |
| 85 | inpc_histologic_group | character | 995 | 100% | 1 | NA |
| 86 | gastric_esophageal_junction_involvement | character | 995 | 100% | 1 | NA |
| 87 | progression_or_recurrence | character | 995 | 100% | 1 | unknown |
| 88 | demographic_id | character | 995 | 100% | 995 | 0059d727-d660-43b7-bfc5-e8e374f813bf, 00aa6c25-74bf-4a10-b015-3873314906b5, 00af936c-0b23-4c49-94ac-254f89600cf0 |
Data Completeness
Missing data rates across clinical variables, highlighting fields with substantial missingness.
83 of 88 variables are fully complete. Only variables with missing data are shown below. See data dictionary for variable definitions.
| Variable | Missing Count | Missing Percent | |
|---|---|---|---|
| 1 | year_of_birth | 995 | 100.0% |
| 2 | year_of_death | 995 | 100.0% |
| 3 | cause_of_death | 807 | 81.1% |
| 4 | days_to_death | 804 | 80.8% |
| 5 | days_to_birth | 42 | 4.2% |
Quality Control
Quality control identifies outlier samples based on library size and gene detection rate. See R/02_quality_control.R for filtering criteria.
The per-sample expression summary (Total Counts, Genes Detected) is shown in the RNA-seq Data section above. This section adds QC-specific metrics: size factors, count dispersion (MAD), and outlier flags.
Variable definitions (see also Glossary):
- Total Counts — library size: sum of all read counts mapped to genes in one sample
- Detected Genes — number of genes with at least 1 mapped read (count > 0)
- Median Count — median read count across all genes in a sample (most genes have 0 or low counts, so this is typically 0)
- MAD Count — median absolute deviation of gene counts within a single sample (measures spread of the count distribution for that sample)
- Size Factor — library size / median library size across all samples (values near 1.0 = typical; <<1 = under-sequenced; >>1 = over-sequenced)
-
Outlier — flagged
Yesif the sample falls in the bottom 5th percentile of either library size OR genes detected
QC Metrics Summary
- Samples assessed: 100
- Outliers flagged: 9
| Sample | Total Counts | Detected Genes | Median Count | MAD Count | Size Factor | Outlier | |
|---|---|---|---|---|---|---|---|
| 1 | MMRF_1618_1_BM_CD138pos | 37,391,891 | 27,609 | 0.0 | 0.0 | 0.644 | Yes |
| 2 | MMRF_2229_1_BM_CD138pos | 51,745,321 | 28,905 | 0.0 | 0.0 | 0.892 | No |
| 3 | MMRF_2143_1_BM_CD138pos | 59,392,637 | 28,249 | 0.0 | 0.0 | 1.023 | No |
| 4 | MMRF_2054_1_BM_CD138pos | 63,680,199 | 32,356 | 1.0 | 1.5 | 1.097 | No |
| 5 | MMRF_1700_2_BM_CD138pos | 96,656,035 | 37,358 | 3.0 | 4.4 | 1.666 | No |
| 6 | MMRF_1871_1_BM_CD138pos | 46,761,239 | 28,751 | 0.0 | 0.0 | 0.806 | No |
| 7 | MMRF_1293_1_BM_CD138pos | 50,232,734 | 30,109 | 0.0 | 0.0 | 0.866 | No |
| 8 | MMRF_1715_1_BM_CD138pos | 58,060,644 | 30,504 | 1.0 | 1.5 | 1.001 | No |
| 9 | MMRF_1637_1_BM_CD138pos | 41,424,544 | 30,874 | 1.0 | 1.5 | 0.714 | No |
| 10 | MMRF_1543_1_BM_CD138pos | 44,701,993 | 27,289 | 0.0 | 0.0 | 0.770 | No |
| 11 | MMRF_1698_1_BM_CD138pos | 46,684,067 | 25,832 | 0.0 | 0.0 | 0.804 | Yes |
| 12 | MMRF_1716_1_BM_CD138pos | 46,903,826 | 27,259 | 0.0 | 0.0 | 0.808 | Yes |
| 13 | MMRF_2341_1_BM_CD138pos | 71,582,628 | 33,440 | 1.0 | 1.5 | 1.234 | No |
| 14 | MMRF_2085_3_BM_CD138pos | 101,561,094 | 34,138 | 1.0 | 1.5 | 1.750 | No |
| 15 | MMRF_2422_1_BM_CD138pos | 49,591,878 | 28,663 | 0.0 | 0.0 | 0.855 | No |
| 16 | MMRF_2429_1_BM_CD138pos | 44,521,218 | 29,397 | 0.0 | 0.0 | 0.767 | No |
| 17 | MMRF_2245_1_BM_CD138pos | 55,815,128 | 30,949 | 1.0 | 1.5 | 0.962 | No |
| 18 | MMRF_2043_1_BM_CD138pos | 70,996,601 | 30,698 | 1.0 | 1.5 | 1.223 | No |
| 19 | MMRF_2401_2_BM_CD138pos | 76,584,241 | 32,974 | 1.0 | 1.5 | 1.320 | No |
| 20 | MMRF_2098_1_BM_CD138pos | 57,538,387 | 29,719 | 0.0 | 0.0 | 0.992 | No |
| 21 | MMRF_2238_1_BM_CD138pos | 66,314,228 | 33,476 | 1.0 | 1.5 | 1.143 | No |
| 22 | MMRF_1436_1_BM_CD138pos | 57,999,952 | 29,968 | 0.0 | 0.0 | 0.999 | No |
| 23 | MMRF_1108_1_BM_CD138pos | 42,289,892 | 28,079 | 0.0 | 0.0 | 0.729 | No |
| 24 | MMRF_1856_1_BM_CD138pos | 48,755,441 | 28,313 | 0.0 | 0.0 | 0.840 | No |
| 25 | MMRF_2746_1_BM_CD138pos | 74,850,398 | 35,919 | 2.0 | 3.0 | 1.290 | No |
| 26 | MMRF_1137_4_BM_CD138pos | 55,587,700 | 32,731 | 1.0 | 1.5 | 0.958 | No |
| 27 | MMRF_1361_1_BM_CD138pos | 76,274,233 | 34,427 | 1.0 | 1.5 | 1.314 | No |
| 28 | MMRF_1048_1_BM_CD138pos | 46,040,142 | 28,127 | 0.0 | 0.0 | 0.793 | No |
| 29 | MMRF_2828_1_BM_CD138pos | 68,349,206 | 27,973 | 0.0 | 0.0 | 1.178 | No |
| 30 | MMRF_2601_1_BM_CD138pos | 39,797,530 | 28,415 | 0.0 | 0.0 | 0.686 | No |
| 31 | MMRF_2232_1_BM_CD138pos | 81,630,421 | 31,401 | 1.0 | 1.5 | 1.407 | No |
| 32 | MMRF_2253_1_BM_CD138pos | 63,958,386 | 32,111 | 1.0 | 1.5 | 1.102 | No |
| 33 | MMRF_1824_1_BM_CD138pos | 37,401,747 | 28,419 | 0.0 | 0.0 | 0.645 | Yes |
| 34 | MMRF_2197_1_BM_CD138pos | 65,930,119 | 33,595 | 1.0 | 1.5 | 1.136 | No |
| 35 | MMRF_2664_1_BM_CD138pos | 64,857,697 | 31,545 | 1.0 | 1.5 | 1.118 | No |
| 36 | MMRF_1819_1_BM_CD138pos | 49,401,746 | 28,052 | 0.0 | 0.0 | 0.851 | No |
| 37 | MMRF_2716_1_BM_CD138pos | 91,426,018 | 34,504 | 1.0 | 1.5 | 1.575 | No |
| 38 | MMRF_1617_1_BM_CD138pos | 37,826,511 | 27,819 | 0.0 | 0.0 | 0.652 | No |
| 39 | MMRF_2199_1_BM_CD138pos | 52,651,580 | 28,971 | 0.0 | 0.0 | 0.907 | No |
| 40 | MMRF_1223_2_BM_CD138pos | 64,253,003 | 30,235 | 0.0 | 0.0 | 1.107 | No |
| 41 | MMRF_1216_1_BM_CD138pos | 54,430,074 | 31,804 | 1.0 | 1.5 | 0.938 | No |
| 42 | MMRF_2705_1_BM_CD138pos | 149,432,793 | 35,625 | 2.0 | 3.0 | 2.575 | No |
| 43 | MMRF_2257_1_BM_CD138pos | 52,847,814 | 32,503 | 1.0 | 1.5 | 0.911 | No |
| 44 | MMRF_2815_1_BM_CD138pos | 54,076,776 | 31,299 | 1.0 | 1.5 | 0.932 | No |
| 45 | MMRF_2636_1_BM_CD138pos | 79,930,092 | 31,067 | 1.0 | 1.5 | 1.377 | No |
| 46 | MMRF_1651_1_BM_CD138pos | 38,531,104 | 29,803 | 0.0 | 0.0 | 0.664 | No |
| 47 | MMRF_1991_1_BM_CD138pos | 69,001,805 | 31,154 | 1.0 | 1.5 | 1.189 | No |
| 48 | MMRF_1502_1_BM_CD138pos | 60,836,136 | 31,923 | 1.0 | 1.5 | 1.048 | No |
| 49 | MMRF_1049_4_BM_CD138pos | 67,752,904 | 31,171 | 1.0 | 1.5 | 1.168 | No |
| 50 | MMRF_1250_1_BM_CD138pos | 41,495,680 | 30,909 | 1.0 | 1.5 | 0.715 | No |
| 51 | MMRF_1496_1_PB_CD138pos | 79,637,573 | 32,560 | 1.0 | 1.5 | 1.372 | No |
| 52 | MMRF_1496_1_BM_CD138pos | 149,615,704 | 34,145 | 1.0 | 1.5 | 2.578 | No |
| 53 | MMRF_1235_1_BM_CD138pos | 60,749,662 | 31,612 | 1.0 | 1.5 | 1.047 | No |
| 54 | MMRF_1978_1_BM_CD138pos | 58,788,425 | 31,621 | 1.0 | 1.5 | 1.013 | No |
| 55 | MMRF_2089_3_BM_CD138pos | 87,760,456 | 34,575 | 1.0 | 1.5 | 1.512 | No |
| 56 | MMRF_1900_1_BM_CD138pos | 52,217,137 | 32,128 | 1.0 | 1.5 | 0.900 | No |
| 57 | MMRF_2224_1_BM_CD138pos | 54,403,435 | 31,139 | 1.0 | 1.5 | 0.938 | No |
| 58 | MMRF_2458_1_BM_CD138pos | 56,294,068 | 29,650 | 0.0 | 0.0 | 0.970 | No |
| 59 | MMRF_1092_1_BM_CD138pos | 36,748,951 | 29,135 | 0.0 | 0.0 | 0.633 | Yes |
| 60 | MMRF_2822_1_BM_CD138pos | 60,054,775 | 32,643 | 1.0 | 1.5 | 1.035 | No |
| 61 | MMRF_1538_1_BM_CD138pos | 61,613,394 | 29,038 | 0.0 | 0.0 | 1.062 | No |
| 62 | MMRF_1992_1_PB_CD138pos | 40,521,003 | 26,760 | 0.0 | 0.0 | 0.698 | Yes |
| 63 | MMRF_2729_1_BM_CD138pos | 94,208,666 | 35,598 | 2.0 | 3.0 | 1.623 | No |
| 64 | MMRF_1625_1_BM_CD138pos | 38,223,103 | 25,966 | 0.0 | 0.0 | 0.659 | Yes |
| 65 | MMRF_2107_1_BM_CD138pos | 63,872,226 | 31,475 | 1.0 | 1.5 | 1.101 | No |
| 66 | MMRF_2055_1_BM_CD138pos | 41,457,997 | 29,706 | 0.0 | 0.0 | 0.714 | No |
| 67 | MMRF_1944_1_BM_CD138pos | 47,103,245 | 30,647 | 1.0 | 1.5 | 0.812 | No |
| 68 | MMRF_1110_2_BM_CD138pos | 77,343,762 | 30,631 | 1.0 | 1.5 | 1.333 | No |
| 69 | MMRF_2228_1_BM_CD138pos | 68,086,550 | 33,922 | 1.0 | 1.5 | 1.173 | No |
| 70 | MMRF_1129_1_BM_CD138pos | 63,899,643 | 31,634 | 1.0 | 1.5 | 1.101 | No |
| 71 | MMRF_1783_2_BM_CD138pos | 82,754,792 | 35,563 | 2.0 | 3.0 | 1.426 | No |
| 72 | MMRF_1783_1_BM_CD138pos | 50,776,425 | 30,821 | 1.0 | 1.5 | 0.875 | No |
| 73 | MMRF_2832_1_BM_CD138pos | 61,360,677 | 28,685 | 0.0 | 0.0 | 1.057 | No |
| 74 | MMRF_1778_1_PB_CD138pos | 67,550,482 | 34,614 | 2.0 | 3.0 | 1.164 | No |
| 75 | MMRF_1267_1_BM_CD138pos | 30,377,770 | 27,628 | 0.0 | 0.0 | 0.523 | Yes |
| 76 | MMRF_2843_1_BM_CD138pos | 77,359,789 | 31,052 | 1.0 | 1.5 | 1.333 | No |
| 77 | MMRF_2047_1_BM_CD138pos | 39,751,015 | 29,294 | 0.0 | 0.0 | 0.685 | No |
| 78 | MMRF_1613_1_BM_CD138pos | 44,241,364 | 29,485 | 0.0 | 0.0 | 0.762 | No |
| 79 | MMRF_1445_1_BM_CD138pos | 53,144,209 | 30,277 | 0.0 | 0.0 | 0.916 | No |
| 80 | MMRF_1556_1_BM_CD138pos | 74,700,167 | 32,015 | 1.0 | 1.5 | 1.287 | No |
| 81 | MMRF_2827_1_BM_CD138pos | 61,996,840 | 32,601 | 1.0 | 1.5 | 1.068 | No |
| 82 | MMRF_1252_1_BM_CD138pos | 22,018,481 | 26,953 | 0.0 | 0.0 | 0.379 | Yes |
| 83 | MMRF_1533_2_BM_CD138pos | 46,280,409 | 30,627 | 1.0 | 1.5 | 0.798 | No |
| 84 | MMRF_1912_1_PB_CD138pos | 55,599,192 | 32,521 | 1.0 | 1.5 | 0.958 | No |
| 85 | MMRF_2153_1_BM_CD138pos | 62,246,722 | 32,147 | 1.0 | 1.5 | 1.073 | No |
| 86 | MMRF_1356_1_BM_CD138pos | 48,785,009 | 30,935 | 1.0 | 1.5 | 0.841 | No |
| 87 | MMRF_1033_1_BM_CD138pos | 52,817,165 | 28,544 | 0.0 | 0.0 | 0.910 | No |
| 88 | MMRF_1079_1_BM_CD138pos | 81,118,832 | 32,913 | 1.0 | 1.5 | 1.398 | No |
| 89 | MMRF_1401_2_BM_CD138pos | 67,018,253 | 27,599 | 0.0 | 0.0 | 1.155 | No |
| 90 | MMRF_1621_1_BM_CD138pos | 46,791,382 | 29,873 | 0.0 | 0.0 | 0.806 | No |
| 91 | MMRF_2225_1_BM_CD138pos | 58,724,738 | 32,595 | 1.0 | 1.5 | 1.012 | No |
| 92 | MMRF_1501_1_BM_CD138pos | 60,234,555 | 32,331 | 1.0 | 1.5 | 1.038 | No |
| 93 | MMRF_2763_1_BM_CD138pos | 48,752,427 | 29,503 | 0.0 | 0.0 | 0.840 | No |
| 94 | MMRF_2150_1_BM_CD138pos | 55,307,603 | 27,982 | 0.0 | 0.0 | 0.953 | No |
| 95 | MMRF_1730_1_BM_CD138pos | 60,872,279 | 29,038 | 0.0 | 0.0 | 1.049 | No |
| 96 | MMRF_2675_1_BM_CD138pos | 73,461,172 | 32,431 | 1.0 | 1.5 | 1.266 | No |
| 97 | MMRF_1690_1_BM_CD138pos | 56,634,991 | 29,767 | 0.0 | 0.0 | 0.976 | No |
| 98 | MMRF_1932_1_BM_CD138pos | 58,691,302 | 32,437 | 1.0 | 1.5 | 1.011 | No |
| 99 | MMRF_2072_1_BM_CD138pos | 78,601,932 | 34,471 | 2.0 | 3.0 | 1.354 | No |
| 100 | MMRF_1462_3_BM_CD138pos | 49,218,987 | 29,169 | 0.0 | 0.0 | 0.848 | No |
After Filtering
- Samples retained: 91
- Genes retained: 30,675
- Genes removed: 29,985 (49.4%)
Data Sources
Results in this vignette are derived from the MMRF CoMMpass study (MMRF-COMMPASS, ~1,143 patients), downloaded via TCGAbiolinks. The pipeline runs with a configurable sample_limit (default 200; CI uses 20).
For full citations, data access tiers, and the distinction between pipeline data and synthetic test data, see the Data Sources vignette.
Recent Changes
Recent project commits with lines added, files changed, and change categories.
| date | type | summary | n_files | lines_added | lines_removed | file_categories |
|---|---|---|---|---|---|---|
| 2026-03-14 | Bug Fix | fix(pipeline): Fix 11 NULL targets — DE condition, ID matching, consensus type | 41 | 146 | 47 | Other, R Source |
| 2026-03-14 | Bug Fix | fix(cachix): Remove –watch-mode auto flag (already default) | 1 | 1 | 1 | Other |
| 2026-03-14 | Bug Fix | fix(pipeline): Fix 3 NULL-target bugs, auto-generate package.nix (#93) | 87 | 235 | 80 | Config, Docs, Other, R Source |
| 2026-03-14 | Bug Fix | fix(nix): Fix cachix signing key, rebuild Bioconductor-dependent targets | 2 | 0 | 0 | Other |
| 2026-03-14 | New Feature | feat(captions): Add dynamic captions to 34 table/plot targets | 22 | 579 | 89 | Other, R Source |
| 2026-03-14 | Bug Fix | fix(vignettes): Enforce zero-computation rule — 22 violations → 0 | 32 | 360 | 764 | Other, R Source, Vignettes |
| 2026-03-13 | Bug Fix | fix(vignettes): Convert kable RDS to data.frames, fix telemetry eval guards | 18 | 8 | 2 | Other, Vignettes |
| 2026-03-13 | Bug Fix | fix(ci): Save data frames (not DT widgets) to RDS for Nix portability | 3 | 0 | 0 | Other |
| 2026-03-13 | Bug Fix | fix(vignettes): Use Quarto #| eval syntax for pkgdown-banner chunks | 11 | 44 | 11 | Vignettes |
| 2026-03-13 | Refactoring | refactor(targets): Move Bioconductor packages to per-target declarations | 11 | 35 | 17 | Other, R Source, Vignettes |
| 2026-03-13 | New Feature | feat(vignettes): Add code provenance, kable→DT conversion, caption compliance | 35 | 1004 | 437 | CI/CD, Other, R Source, Vignettes |
| 2026-03-13 | Bug Fix | fix(vignettes): Skip NULL RDS in safe_tar_read, return invisible(NULL) | 11 | 22 | 22 | Vignettes |
| 2026-03-13 | Bug Fix | fix(glossary): Prevent double DT::datatable() wrapping in glossary-table chunk | 1 | 3 | 1 | Vignettes |
| 2026-03-13 | CI/CD | ci: Show quarto errors with quiet=FALSE, render individual vignettes in diagnostic | 1 | 20 | 6 | CI/CD |
| 2026-03-13 | CI/CD | ci: Add verbose quarto error diagnostics on build failure | 1 | 14 | 1 | CI/CD |
| 2026-03-13 | Bug Fix | fix(vignettes): Strip Nix paths from DT widgets, auto-wrap data frames | 25 | 66 | 28 | CI/CD, Other, Vignettes |
| 2026-03-13 | CI/CD | ci: Add diagnostic quarto render step to debug build failure | 1 | 17 | 0 | CI/CD |
| 2026-03-13 | Bug Fix | fix(vignettes): Revert safe_tar_read placeholder, guard gene-report | 11 | 12 | 56 | Vignettes |
| 2026-03-13 | Maintenance | chore: Export vig_count_distribution_plot as ggplot RDS (513KB) | 1 | 0 | 0 | Other |
| 2026-03-13 | Bug Fix | fix(vignettes): Enable code eval in CI with RDS fallback | 80 | 113 | 74 | CI/CD, Other, R Source, Vignettes |
Reproducibility
Git Commit Info (click to expand)
Generating code
{
if (!requireNamespace("gert", quietly = TRUE))
return(NULL)
tryCatch({
info <- gert::git_info()
log1 <- gert::git_log(max = 1)
git_df <- data.frame(Item = c("Commit Hash", "Author",
"Time", "Branch"), Value = c(info$commit, log1$author,
as.character(log1$time), info$shorthand), stringsAsFactors = FALSE)
DT::datatable(git_df, rownames = FALSE, options = list(pageLength = 10,
dom = "t", scrollX = TRUE), caption = htmltools::tags$caption(style = "caption-side: top; text-align: left;",
"Git repository state at pipeline build time."))
}, error = function(e) NULL)
}| Item | Value |
|---|---|
| Commit Hash | c20a60ac5d4cd228607ef8c3af571c91866a902d |
| Author | John Gavin <john.b.gavin@gmail.com> |
| Time | 2026-03-13 20:37:26 |
| Branch | main |
Session Info (click to expand)
Show code
sessionInfo()
#> R version 4.5.3 (2026-03-11)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.3 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> loaded via a namespace (and not attached):
#> [1] base64url_1.4 gtable_0.3.6 jsonlite_2.0.0
#> [4] dplyr_1.2.0 compiler_4.5.3 tidyselect_1.2.1
#> [7] callr_3.7.6 scales_1.4.0 yaml_2.3.12
#> [10] fastmap_1.2.0 ggplot2_4.0.2 R6_2.6.1
#> [13] generics_0.1.4 igraph_2.2.2 knitr_1.51
#> [16] backports_1.5.0 targets_1.12.0 tibble_3.3.1
#> [19] pillar_1.11.1 RColorBrewer_1.1-3 rlang_1.1.7
#> [22] xfun_0.57 S7_0.2.1 otel_0.2.0
#> [25] cli_3.6.5 withr_3.0.2 magrittr_2.0.4
#> [28] ps_1.9.1 digest_0.6.39 grid_4.5.3
#> [31] processx_3.8.6 secretbase_1.2.0 lifecycle_1.0.5
#> [34] prettyunits_1.2.0 vctrs_0.7.2 evaluate_1.0.5
#> [37] glue_1.8.0 data.table_1.18.2.1 farver_2.1.2
#> [40] codetools_0.2-20 rmarkdown_2.30 tools_4.5.3
#> [43] pkgconfig_2.0.3 htmltools_0.5.9